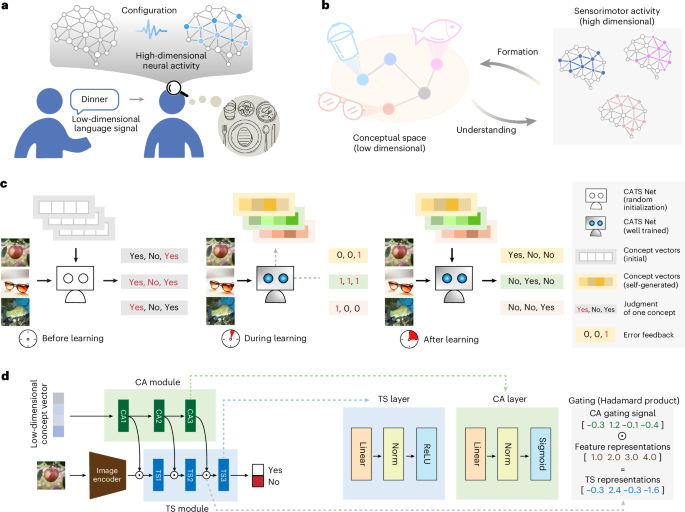
On convex decision regions in deep network representations
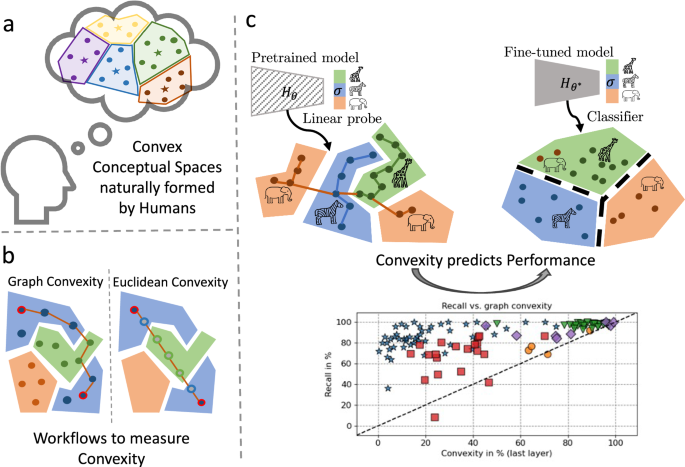
Properties of euclidean convex sets
A set in Euclidean spaces is convex if for every two points from this set, the segment (i.e. shortest path, denoted y(t) with t ∈ [0, 1]), between these points is also within the set. Formally:
Definition 1
(Euclidean convexity). A subset \(S\subset {{\mathbb{R}}}^{D}\) is convex iff ∀ x, y ∈ S ∀ t ∈ [0, 1], y(t)= tx + (1 − t)y is also in S39.
From the definition, it follows that the intersection of two convex sets is also a convex set. Hence, conceptual conjunction (AND operation) preserves convexity. Disjunctions (OR operations), however, do not, since the union of convex sets is not necessarily convex (it is trivial to construct counter-examples)39. Euclidean convexity is conserved under affine transformations, and hence convexity is robust to relevant latent space transformations in deep networks (see a formal proof in Supp. Mat. 1.2). Euclidean convexity is closely related to conjunctions of linear classifiers. In fact, a convex set can alternatively be defined as the intersection of linear half-spaces (possibly infinite), e.g. implemented by a set of linear decision functions resulting in a polyhedron39.
The relevant geometric structure of deep networks is not necessarily Euclidean, hence, we will also investigate the convexity of decision regions in data manifolds with more general geometry – Riemannian manifolds. The generalization of a segment to Riemannian manifolds is a geodesic, hence, we need to use geodesics as shortest paths between two points on a manifold. Formally, in a Riemannian manifold M with metric tensor g, the length L of a continuously differentiable curve y: [0, 1] → M is defined by \(L({{{\bf{y}}}})=\int_{0}^{1}\sqrt{{g}_{{{{\bf{y}}}}(t)}(\dot{{{{\bf{y}}}}}(t),\dot{{{{\bf{y}}}}}(t))}dt\), where \(\dot{{{{\bf{y}}}}}(t):=\frac{\partial }{\partial t}{{{\bf{y}}}}(t)\). A geodesic from x to y is a curve connecting y(0) = x and y(1) = y, minimizing this length, i.e. \({{{\rm{geodesic}}}}({{{\bf{x}}}},{{{\bf{y}}}})={\arg \min }_{{{{\bf{y}}}}}L({{{\bf{y}}}})\). While geodesics are unique for Euclidean spaces, they may not be unique in manifolds. We can now generalize to geodesic convexity in manifolds:
Definition 2
(Geodesic convexity). A region S ∈ M is geodesic convex, iff ∀ x, y ∈ S, there exists at least one geodesic y(t) connecting x and y, that is entirely contained in S.
When modeling latent spaces with sampled data, we must further transform the above definitions to data-driven estimators, such efforts are reported, e.g. in14,15. In this work, we choose a simple approach inspired by ISOMAP18, hence based on graph convexity in data manifolds.
Definition 3
(Graph convexity, see e.g.40). Let (V, E) be a graph and A ⊆ V. We say that A is convex if for all pairs x, y ∈ A, there exists a shortest path P = (x = v0, v1, v2, …, vn−1, y = vn) and ∀ i ∈ {0, …, n}: vi ∈ A.
For reasonably sampled data, we can form the graph based on Euclidean nearest neighbors (as in ISOMAP), effectively creating an undirected, weighted graph. The shortest path between two points within such a graph is defined as the path, which minimizes the sum of weights of the edges41.
We note two important properties of this estimator, first, the graph-based approximate convexity measure is invariant to isometric transformation and uniform scaling (see a formal proof in Supp. Mat. 1.3), and second, the sample-based estimator of convexity is consistent.
As we will measure convexity in labeled subgraphs within larger graphs, Dijkstra’s algorithm is preferred over the Floyd-Warshall algorithm used in ISOMAP. Dijkstra’s algorithm finds the shortest path from a given node to each of the other N nodes in the graph with E edges in \({{{\mathcal{O}}}}(N\log N+E)\)41,42, while Floyd-Warshall efficiently finds the shortest distance between all vertices in the graph in \({{{\mathcal{O}}}}({N}^{3})\)43,44. As we have a sparse graph with E ≪ N2, Dijkstra’s algorithm will be more efficient. With these approximations, we are in a position to create a graph-based workflow for quantifying convexity in Euclidean and manifold-based structures. Note that for sampled data, we expect a certain level of noise, and hence convexity will be graded.
The consistency of sample/graph-based geodesic estimates has been discussed in connection with the introduction of ISOMAP45 and more generally in46. Graph connectivity-based estimates of geodesics from sample data are implemented using two procedures: The neighborhood graph can be determined by a distance cutoff ϵ, so that any points within the Euclidean distance ϵ are considered neighbors, or by K-nearest neighbors (KNN) based on the Euclidean distance. Consistency of these estimates, i.e., that the sample-based shortest paths converge to geodesics, is most straightforward to prove for the former approach45,46. The consistency proof for the ϵ-based procedure is based on the smoothness of the metric, a uniformly bounded data distribution (1/c < p(x) < c) and scaling of the distance cutoff or K so the connectivity (number of edges per node) increases for large samples (cutoff decays slowly ϵ → 0 as sample size N → ∞)46.
In finite samples, a (too) large connectivity will bias the geodesics, while a (too) small connectivity can lead to disconnected graphs and noisy estimates. We explore the role of the way we construct the graph. The consistency proof holds for a graph constructed with a distance cutoff ϵ. We compare it to graphs constructed by keeping K-nearest neighbors and symmetrizing the graph. We chose the number of nearest neighbors to be 3, 5, 10 and 20. The respective ϵ values were chosen to keep approximately the same number of edges in the graph.
Figure 2 d shows that the graph constructed with a distance cutoff ϵ is very disconnected, and the scores computed with this type of neighborhood are biased towards zero (see Fig. 2e). If we skip the disconnected pairs and compute the score only from existing paths, the results are very close to the scores based on K-nearest neighbors (see Fig. 2f). All experiments also demonstrate that the role of the size of K is negligible. Therefore, we choose a KNN-based approach with K = 10 in the complete set of experiments.

a Graph convexity can be more general than Euclidean convexity for a sufficiently large number of data points, N, and an appropriate number of nearest neighbors, K. An Euclidean convex set can have a graph convexity score lower than 1 in the case of isolated subgraphs of one class, connected only through the other class. b Smaller values of K can lead to a disconnected graph with a low graph convexity score due to missing paths between the nodes from the same class. Small N may lead to edges to distant points that are then used in the shortest paths. c Difference between computing the graph convexity scores of the data labels vs. the model labels in the last layer. Model labels create Euclidean-convex regions (left), which is not the case for data labels (right) if the model’s accuracy is lower than 100%. d The graph constructed with a distance cutoff ϵ is very disconnected, resulting in many paths that don’t exist, i.e. data points that cannot be connected via neighbors. The graph constructed using kNN is more complete, leading to all data points being able to connect via a path through their neighbors, independent of K. e The graph convexity scores computed with disconnected neighborhoods (based on ϵ) are biased towards zero, while the kNN-based graph leads to stable convexity scores, again independent of K. f If we skip the disconnected pairs and compute the score only from existing paths, the results of all graph construction methods are very similar. The results in (d)–(f) are computed on 100 ImageNet classes with data2vec.
Neural networks and convexity
Should we expect convexity of decision regions of neural network representations? In fact, several mechanisms could contribute to the promotion of convexity. First, the ubiquitous softmax is essentially a convexity-inducing device, hence, typical classification heads will induce convexity in their pre-synaptic activation layer. This is most easily seen by noting that softmax decision regions (maximum posterior decisions) are identical to the decision regions of a linear model, and linear models implement convex decision regions (see Supp. Mat. 1.1). Secondly, many recent models are based on transformer architectures with attention heads47. These heads contain softmax functions and thus induce convexity in their weighing of attention. Thirdly, typical individual artificial neurons, e.g. ReLUs, are latent half-space detectors, and half-spaces are convex as noted above. Note that deep multi-layer perceptrons can approximate any non-convex decision region, including disconnected decision regions48. These findings suggest that convexity is possible in neural networks, although it is not forced by design.
An extensive body of work concerns the geometry of input space representations in ReLU networks and has led to a detailed understanding of the role of so-called ‘linear regions’: In networks based on ReLU nonlinearity, the network’s output is a piecewise linear function over convex input space polyhedra, formed by the intersection of neuron half-spaces49,50,51,52. Interestingly, in typical networks (e.g. at initialization or after training), the linear regions are much less in number and simpler in structure compared to theoretical upper bounds50,51,52. During training, the collection of convex linear regions is transformed and combined into decision regions through the later network layers. The resulting decision regions are, therefore, unions of convex sets and may in general be non-convex or non-connected, as noted in ref. 48. Our two measures of convexity, Euclidean and graph-based, probe different mechanisms of generalization, the former is associated with generalization by linear interpolation, while the latter is associated with generalization by interpolation along the data manifolds. As both mechanisms may be relevant for explaining generalization in cognitive systems, we are interested in the abundance and potential role of both measures of convexity. Note that a region may be convex in terms of the Euclidean measure but not the graph-based (e.g. a decision region formed by disconnected but linearly separated subsets) and a region may be graph-convex, but not Euclidean-convex (e.g., a general shaped connected region in the data manifold). For visual examples, see Fig. 2a.
Convexity measurement workflows
We developed and presented two workflows for measuring graph convexity and Euclidean convexity respectively in latent spaces of neural networks.
We are interested in measuring the approximate convexity of a decision region, here, a subset of nodes in a graph. A decision region for a specific class is a set of points that the model classifies as belonging to this class. We measure convexity per layer and per class separately.
We first create a graph containing the representations of the data points of all classes. The points are nodes in the graph and the Euclidean distances between the nodes are the weights of the edges. To handle manifold-based representation, for each node, we create an undirected edge only to the nearest neighbors (K = 10). This procedure creates a sparse undirected weighted graph with positive weights (i.e. distances) only.
We now sample Ns pairs of points within the given predicted class label and compute the shortest path in the graph between the pairs using Dijkstra’s algorithm41. For each path, we compute a score between 0 and 1. The path score is defined as the proportion of the number of nodes on the shortest path, without the endpoints, inside the decision region (i.e. with the same predicted label). If an edge directly connects the pair of nodes, the score is 1. If the points are not connected, the score is 0. We average the scores for all paths within one class to get a convexity score per class per layer. To get one convexity score per layer, we average over all paths regardless of class. Uncertainties are calculated as the standard error of the mean, where we set n as the number of points in the given class. This is likely a conservative estimate since the mean is based on many more pairs than there are points.
The runtime complexity of this procedure is \({{{\mathcal{O}}}}({N}^{2}(D+1)+{N}_{c}{N}_{s}N\left(\log N+K\right))\), where N denotes the number of data points, D is the dimensionality of the representations and Nc is the number of classes. See Supp. Mat. 1.4 for details.
Euclidean convexity is also measured for each layer and class separately. To measure the convexity of class C in layer l, we first extract hidden representations at l of all points predicted to belong to class C. We sample Ns pairs of points. For each pair, we compute the Np = 10 equidistant points on the linear segment connecting the representations of the two endpoints. We feed the interpolated points to the rest of the network (after layer l). The score for each pair is then the proportion of the interpolated points that are also predicted to belong to class C. Finally, to get the score of Euclidean convexity for the whole class, we average the scores over all the pairs of points. To get the Euclidean convexity for each layer, we average over all classes.
The runtime complexity of this procedure is \({{{\mathcal{O}}}}({N}_{c}{N}_{s}{N}_{p}D)\), where Nc denotes the number of classes, and D is the dimensionality of the representations. See Supp. Mat. 1.4 for details.
To gain more intuition about the two types of convexity and their differences, we present examples of various properties of the graph and Euclidean convexity score on synthetic data. Figure 2a showcases two examples of data distributions that are either graph or Euclidean convex but not the other, this shows that graph convexity can be more general than Euclidean convexity and capture more complex structures. Figure 2b shows how different values of the sampled data (N) and nearest neighbors (K) influence the created graph and, therefore, also the graph convexity score. A high enough N and K is needed to guarantee a stable result. For too small N or K, the graph convexity score deteriorates and fails to provide a good estimate of the convexity in the data structures. In experiments (see Fig. 2d–f) we found that the influence of K is negligible given an appropriate N. Another consideration is the type of class labels, where we have two options: data labels (true classes) and model labels determined by the predictions of a model (decision regions). Figure 2c illustrates the difference between these two in the last layer of a model. From Theorem 3 in Supp. Mat. 1.3, we have that the last layer is always Euclidean convex for model labels. We focus on decision regions defined by model labels as they directly probe model representations and decision processes. Note that Euclidean convexity is always based on model labels since it is based on the prediction of interpolated points, while graph convexity can be computed for data as well as model labels.
For the pretrained models, we obtain the predicted labels by training a linear layer on top of the last hidden layer (with the rest of the model frozen). This procedure is similar to linear-classifier-based probes that are widely used in natural language processing to understand the presence of concepts in latent spaces53,54. A prominent example of probe-based explanation in image classification is the CAV (concept activation vector) scheme proposed by Kim et al.55, in which auxiliary labeled data sets are used to identify concept directions with linear classifiers (concept class versus random data). In our approach, we use a multi-label probe of the last layer to identify the model labels and use these labels to compute and compare convexity throughout the network.
The score depends on the number of classes, and also on the number of predicted data points per class. It follows that the exact numbers are not directly comparable across modalities. A scale for convexity can be set using a null hypothesis that there is no relation. Under this null, we can estimate convexity with randomized class assignments and get a baseline score. For balanced datasets we find a baseline score of 1/C with C being the number of classes.
Measurements based on neighbors in high-dimensional data can be sensitive to the so-called hubness problem56. We evaluated the hubness of the latent representations in terms of k-skewness and the Robinhood score. Results are deferred to Supp. Mat., Section 2.1, since only mild hubness issues were detected for most domains. We decided to analyze convexity without adjustment for hubness, to avoid possible biases introduced by normalization schemes56.
Convexity of latent representations before and after fine-tuning
To gain intuition on how and where convex regions form and the effect of fine-tuning on this process, we first inspect the t-SNE plots, which are a projection of high-dimensional data into 2D57. The t-SNE plots for a subset of layers of all modalities can be seen in Fig. 3a. From Theorem 3 in Supp. Mat. 1.3, we know that the last layer of both the pretrained and fine-tuned models is Euclidean convex. The t-SNE plot illustrates this quite clearly for the fine-tuned model, where the classes are clearly clustered and separated in the last layer. The picture is not quite as evident for the pretrained model, however, even here we see a clustering of the classes. We also see that points with the same predictions get clustered already within earlier layers. The observed structure in these low-dimensional representations adds motivation to our investigation of the convexity of class labels using Euclidean and graph methods in the high-dimensional latent spaces.
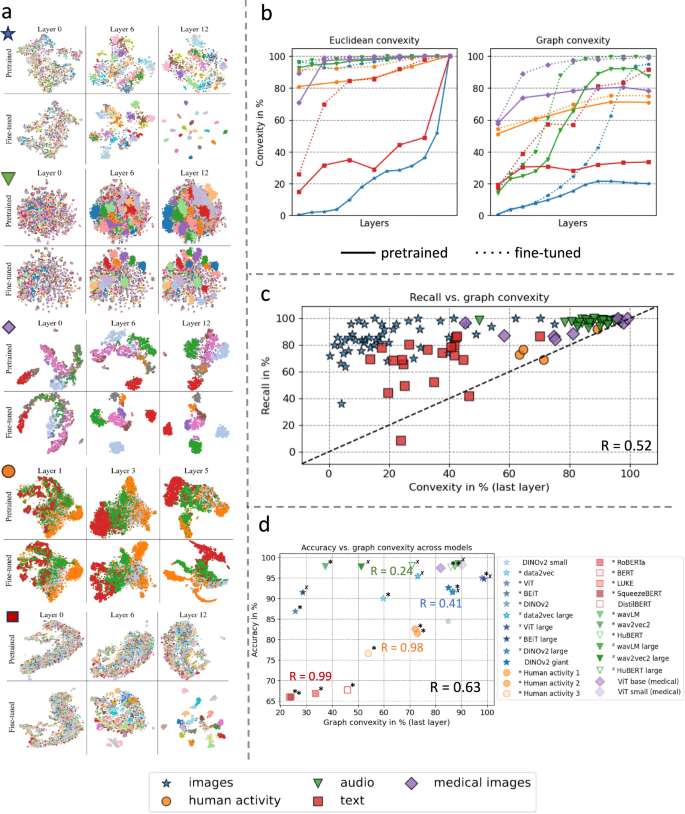
a t-SNE plots show the forming of clustered regions already in pretrained models, which develop into more defined and clustered regions during fine-tuning. This gives an intuition of how convex regions form. b Euclidean and graph convexity scores for all modalities for decision regions of pretrained and fine-tuned networks. Decision regions are determined by model-predicted labels. Fine-tuning generally increases convexity scores. Note that the results are not directly comparable across modalities, as the convexity score depends on the number of classes. c Graph convexity of a subset of classes in the pretrained models (last layer) vs. recall rate of these individual classes in the fine-tuned models for all data domains. We find a positive correlation between the convexity in pretrained models and downstream performance in fine-tuned models. d Graph convexity (last layer) of pretrained models vs. accuracy of fine-tuned models for several models in each data domain. A positive linear relation can be observed for all domains. Models denoted with */× have the same architecture and size within each modality, showing the relation between convexity and performance is not driven by architectural differences.
We investigate how convex regions develop across layers in the networks and the effect of fine-tuning on these decision regions. We find that convexity is pervasive both before and after fine-tuning, while convexity generally increases across layers within a model, and fine-tuning increases convexity. Figure 3b shows the results for all modalities. It is important to note that the magnitudes of the convexity scores differ among modalities because of different amounts of data and numbers of classes. However, the trend is the same for all modalities. Random labeling yields graph convexity scores of approximately \(\frac{1}{C}\), where C is the number of classes. All the convexity scores are significantly higher than these baselines. Generally, class convexity is increased by fine-tuning. Note that the graph convexity in the last layer is (in some cases considerably) lower than 1 ~ 100%, even though this layer is always Euclidean convex. For detailed results of the analysis of individual data modalities including uncertainties, see Supp. Mat. 2.1, Supp. Figure 2.
Convexity and performance
To test the hypothesis motivated by cognitive science that convexity facilitates few-shot learning, we plot the post-finetuning accuracy (recall) per class versus the pretraining convexity scores in the last layer as shown in Fig. 3c for graph-based convexity. There is a strong association between both types of convexity measured in the pretrained model and the accuracy of the given class in the fine-tuned model. Indeed, there is a significant correlation in both cases. The Pearson correlation coefficient between Euclidean convexity and recall is 0.53 ± 0.06 and 0.52 ± 0.06 for graph convexity. In Supp. Mat. 2.5, we explore further the relation between graph convexity and recall, and find that the relation is best modeled by a linear relation.
We also perform experiments comparing multiple models within the same domain. We focus on different models in the image, text, and audio domains, and one model architecture pretrained on various datasets in the human activity domain. Note here that in order to include more models for the image domain, we use a challenging subset of ImageNet with 10 classes to evaluate and fine-tune the models. Figure 3d demonstrates a strong positive correlation (r = 0.63, p = 5e − 4) between graph convexity in the last layer and downstream accuracy. It is important to note that the convexity scores are not directly comparable across domains but only within each domain due to different amounts of classes. Therefore, we report correlations separately for each domain (see Fig. 3d). The positive correlation between convexity and performance is particularly notable in the text, human activity and medical image domain. In the audio domain, the relationship is ambiguous since all models achieve very high accuracy. Generally, we observe that high convexity in pretrained models can be related to good performance in fine-tuned models and suggest that convexity can be used as an indicator for model choice. Ansuini et al. showed that the so-called intrinsic dimension of the last layer in a pretrained model, also predicts downstream performance down58. In Supp. Mat. 2.2 we compare the predictions by convexity and different versions of the intrinsic dimension and we find stronger association for convexity.
link