Dissecting the enhancer gene regulatory network in early Drosophila spermatogenesis
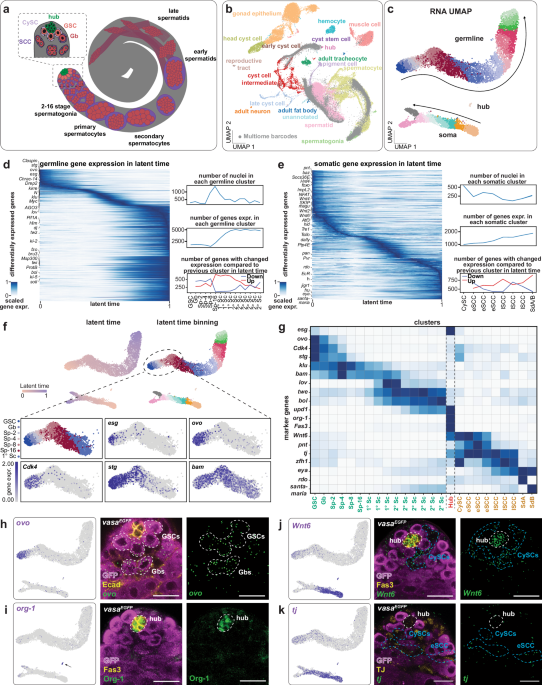
Multimodal profiling reveals rare cells in Drosophila tests
To investigate regulatory programs in the Drosophila testis, we generated a single-nucleus multi-omics atlas by combining RNA expression and chromatin accessibility (snRNA-seq and snATAC-seq) from cells at the apical tip of adult testes. This region contains the stem cell niche, which is essential for understanding stem cell regulation and early differentiation (Fig. 1a and Supplementary Fig. 1a). To enrich underrepresented niche cells in existing single-cell datasets9,11,28, we microdissected tissue, isolated nuclei, and used the 10x Genomics Multiome platform to jointly profile gene expression and chromatin accessibility per nucleus29. We profiled 10,335 high-quality nuclei across four replicates, capturing 13,034 protein-coding genes, 5839 non-coding RNAs, and 46,619 accessible regions. Compared to the Fly Cell Atlas, our dataset is enriched for early germline and somatic cells and depleted of terminally differentiated types, which comprise ~40% of FCA testis data (Fig. 1b and Supplementary Fig. 1d). Transcriptome-based UMAP embedding revealed clear separation of three cell groups: 7856 germline cells, 2428 somatic cells, and 51 hub cells (Fig. 1c). Pseudotime inference and differential gene expression analysis (log2FC > 1.5 for germline, >3 for soma) revealed developmental progression consistent with previous studies9,11. In the germline, late spermatogonia undergo a major transcriptional shift, marked by increased gene expression and a transition to a more transcriptionally active state (Fig. 1d). Gene Ontology (GO) analysis of differentially expressed genes (DEGs) reveals a transition from signaling functions to biosynthesis and remodeling (Supplementary Fig. 1e), consistent with entry into the primary spermatocyte stage. In contrast, somatic differentiation is accompanied by a steady increase in gene expression (Fig. 1e, center right), which stands in contrast to the general trend of transcriptional downregulation during differentiation30. The most pronounced transcriptional changes occur at two inflection points – early and late somatic cyst cells (eSCCs and lSCCs) – characterized by widespread downregulation of genes relative to earlier stages (Fig. 1e, bottom right). GO analysis of somatic DEGs revealed enrichment for metabolic functions (Supplementary Fig. 1f), consistent with the role of somatic cells in supporting germline development31,32.
To refine cell state classification and explore soma-germline coordination, we analyzed transcriptional kinetics using scVelo, which infers latent time from splicing dynamics33. Because germline and somatic cells differ in their transcriptomes and differentiation trajectories, splicing dynamics were computed separately. Cells were binned by stage, like GSCs, gonialblasts (Gbs), 2-cell to 16-cell spermatogonia (Sp-2 to Sp-16), excluding the 51 quiescent hub cells (Fig. 1f). Although no single marker gene was specific to early germline stages, combinations of escargot (esg), ovo, cyclin dependent kinase 4 (cdk4) and string (stg) enabled high-resolution classification (Fig. 1g). For example, the Snail-like TF gene esg, a stem cell marker in other systems34 and known to control CySC maintenance35, is expressed in GSCs, CySCs, and hub cells (Fig. 1f, g). ovo, a zinc-finger TF gene required throughout oocyte maturation36,37, peaks in GSCs and Gbs (Fig. 1g, h), and declines in cells with increasing transcript levels of the differentiation factor encoding gene bag-of-marbles (bam)38 (Fig. 1f, g). The expression of cdk4 in mitotically dividing spermatogonia persists until the peak expression of bam (4/8-cell spermatogonia, Sp-4/8)39 protein-tyrosine-phosphatase encoding gene stg remains in all mitotically dividing cell stages (Fig. 1f, g) up to stage-16 spermatogonia (Sp-16), where the cells undergo four rounds of transit-amplifying mitotic divisions. We validated marker expression patterns in vivo by single-molecule fluorescence in situ hybridization (smFISH)40. As predicted from transcriptomic data, ovo is largely restricted to GSCs and Gbs, despite earlier assumptions of ovary specificity41,42. We also identified org-1 as a novel marker of hub identity (Fig. 1i), and verified expression of known (traffic jam) and novel (DWnt6) somatic markers (Fig. 1j, k).
Together, this dataset resolves rare early populations and enables precise discrimination of closely related cell states in both lineages, in particular in the germline. It recapitulates and extends prior transcriptomic findings and establishes marker gene combinations to molecularly define GSCs, Gbs, and early progenitor cells.
Regulatory region features in Drosophila testis
To align chromatin accessibility with transcriptional dynamics, we leveraged shared barcodes from the 10x Multiome platform to transfer latent time assignments from snRNA-seq to snATAC-seq data, bypassing gene activity imputation and enabling a unified developmental trajectory (Fig. 2a). This integration facilitated downstream analyses of chromatin accessibility, predictive eRegulon inference, and germline-soma signaling predictions.

a UMAP embedding of single-nucleus ATAC-seq data, colored by transcriptome-based latent time clusters, shows distinct accessibility trajectories for germline and soma. (arrows). b Heatmap of label transfer scores between RNA and ATAC datasets. c Left: Representative cluster-specific accessible regions; underlined regions are cluster-enriched. Right: total and cluster-specific accessible regions per cluster. d Left: Distribution of accessible regions by genomic annotation; solid and dashed lines represent germline and somatic regions, respectively. Right: distribution of gene-associated peaks per gene. e, f Accessibility at the zfh1 and stg loci ±20 kb, with cluster-specific tracks and temporal gene expression (insets). g, h Motif enrichment scores (normalized enrichment score, NES) across latent time for population-specific TFs in germline (g) and soma (h), identified via SCENIC+; logos indicate enriched motifs.
We defined 25 transcriptionally distinct clusters and identified 29,989 non-overlapping accessible chromatin peaks using MACS243, composing of 46,619 consensus-accessible regions (500 bp each) as calculated by pycisTopic44, capturing chromatin dynamics across germline and somatic lineages. As expected from the extensive DNA remodeling characteristic of germline development45, accessibility was higher in the germline (34,837 vs. 29,085 regions) compared to the soma, peaking in stage-16 spermatogonia (Sp-16) before declining with spermatocyte differentiation (Fig. 2c). In contrast, accessibility increased in the somatic lineage accompanying the secondary spermatocyte (2°Sc) stage (Fig. 2c), mirroring transcriptomic activity (Fig. 1d, e). This may indicate a shift in the regulatory interplay between germline and soma, coinciding with increased expression of metabolic genes (Supplementary Fig. 1f). Indeed, transcriptome data indicate higher transcriptional activity in somatic cells downstream of the branching point relative to germline cells at comparable stages9,11.
Promoter regions (−1 kb to +100 bp from TSS) accounted for most accessibility44, particularly in spermatogonia and primary spermatocytes (1 °Sc), while intronic accessibility was higher in soma (Fig. 2d). The distribution across exonic, intronic, promoter, and intergenic loci was broadly similar between lineages, with low intergenic representation reflecting the compact Drosophila genome. Most genes were associated with multiple accessible regions by correlative analysis of SCENIC+44 (Fig. 2d, see “Methods”), consistent with the presence of putative cis-regulatory elements, such as enhancers, contributing to gene regulation27,46. Promoter accessibility correlated with gene expression: for example, zfh1, a CySC marker essential for CySC maintenance and non-autonomous GSC support47, showed high TSS accessibility in CySCs, which declined during differentiation alongside transcript levels (Fig. 2e). A similar pattern was observed for stg in the germline (Fig. 2f). Notably, zfh1 accessibility declined more slowly than Zfh1 protein levels, suggesting that while Zfh1 protein becomes restricted to CySCs47, the broader chromatin accessibility landscape persists transiently, potentially reflecting residual transcriptional competence in early differentiating cells.
To assess how chromatin state influences TF activity, we integrated expression with motif enrichment using pycisTarget44, restricting analysis to accessible regions to increase specificity48. This revealed lineage- and stage-specific TF motif availability. For example, the Ovo motif was enriched in spermatogonia (Fig. 2g), implicating Ovo in germline regulation. Similarly, motifs for Mad, a Dpp/BMP effector required for GSC maintenance6, and Klu, a regulator of progenitor commitment in the gut49, were enriched in early germline cells. Interestingly, while Mad and klu transcripts remained high (Fig. 1g), motif enrichment declined, suggesting that chromatin closure may limit their regulatory activity during differentiation. Temporal motif analysis revealed dynamic TF motif accessibility across spermatogenesis. Motifs for Fru, Bab1, and Lov were enriched in primary spermatocytes (1 °Sc), while Hth, Achi, and Vis were enriched in secondary spermatocytes (2 °Sc) (Fig. 2g). Several of these TFs are implicated in germline differentiation21,50,51,52, while others remain uncharacterized. In soma, motif enrichment changed more gradually (Fig. 2h): motifs for Stat92E, Tj, and Bowl—key regulators of early cyst cell development5,53,54—were enriched early and declined with differentiation.
In summary, by linking TF motifs to accessible regions within single nuclei, we reveal how regulatory potential shifts over time, consistent with stage-specific transcriptional programs and lineage progression. These findings would not have been possible by investigating the transcriptomics alone.
eRegulons reveal cell-specific transcription regulators
To uncover the regulatory architecture of spermatogenesis, we integrated snRNA-seq and snATAC-seq data using the SCENIC+ framework44 to construct a testis-specific enhancer gene regulatory network (eGRN). This network comprises modular eRegulons, each linking a TF to correlated accessible regions and predicted target genes at defined cell-stages (Fig. 3a and Supplementary Fig. 2a). TF-gene associations were inferred by expression correlation, refined by accessible TF motif presence within ±50 kb of targets. An illustrative example is the ovo eRegulon (Fig. 3b).

a Schematic of SCENIC+-based enhancer gene regulatory network (eGRN) inference. Regulatory links are defined by correlated TF expression, motif accessibility, and target gene expression. b Example ovo eRegulon with cell stage-specific TF-peak-gene associations in early germline clusters; colors indicate cluster-specific regulatory activity. c Dotplot heatmap showing TF expression (color) and correlation to accessible targets (dot size) across cell types. Lineage-specific eRegulons are ordered by maximal TF expression. See also Supplementary Fig. 2.
Although TF function is influenced by chromatin context and cofactors, most TF-DNA motifs are derived from in vitro data and lack context specificity. To address this, we integrated lineage-resolved differentially accessible regions (DARs), DEGs, and latent time to infer dynamic regulatory activity, revealing stage-specific targets of eRegulons such as ovo in GSCs, Gbs, and 2-cell spermatogonia (Sp-2) clusters (Fig. 3b, c). We identified 103 high-confidence activator eRegulons (Fig. 3c and Supplementary Fig. 2a) and 44 predicted repressors (Supplementary Fig. 2b, c). Activators grouped into germline-, soma-, or dual-lineage classes, while repressors were mostly lineage-specific, with fru as the sole exception, active in both somatic cells and 2°Scs. Several regulatory modules control transcriptional programs at specific stages: in early spermatogonia, ovo, Chrac-14, and klu eRegulons are active, suggesting previously unrecognized roles in early germline regulation. Notably, ovo also functions as a repressor in these cells (Supplementary Fig. 2b, c), consistent with its dual activity in the female germline55. As cells differentiate into primary and secondary spermatocytes (1°Scs, 2°Scs), eRegulon diversity increases. GO analysis using g:Profiler revealed enrichment for chromatin remodelers (Chrac-14, BEAF-32, E(bx), egg, nej, Clamp, Lam, pho) and neuronal regulators (pros, pdm3, nub, optix, Lim3, acj6, ab, ham, Dref), consistent with known testis-brain parallels56. We also identified achi and vis eRegulons, likely mediating recruitment of tMAC components Always early (aly) and Cookie monster (comr) to activate genes required for meiotic progression21. In the somatic lineage, eRegulons primarily regulate signaling. Key TFs include foxo, pointed (pnt), pebbled (peb), Eip75B, cubitus interruptus (ci), and anterior open (aop). For example, pnt negatively regulates EGFR signaling during dorsal follicle cell patterning57, and ci acts as the canonical Hh effector required for CySC self-renewal8. In hub cells, we detected a small set of highly specific eRegulons centered on abd-A, foxo, ci, mirror (mirr), and tj (Supplementary Fig. 2b, c).
In summary, we identified context-specific regulators of spermatogenesis, with germline eRegulons enriched for chromatin and meiotic functions, and somatic modules for signaling pathways. All eRegulons are accessible via our Atlas Web Application (dAWA).
Validating core eRegulon TFs in Drosophila testis
To functionally validate TFs predicted from eRegulon analysis, we performed cell type-specific CRISPR perturbations using the GAL4 > UAS system. All experiments were restricted to adulthood by shifting flies from 18 °C to 29 °C at eclosion to assess the role of the TFs during tissue homeostasis, age-matched controls were treated the same way (Fig. 4a). Activators were prioritized due to the higher false-positive rate among predicted repressors48. Lineage-specific drivers included nos-GAL4,UAS-Cas9 for early germline interference58 and patched (ptc)–GAL4,UAS-Cas9 for early somatic lineages3 (Supplementary Fig. 3a). TF depletion efficiency after CRISPR was confirmed via smFISH quantification of transcript levels in adult testes. Of nine TFs tested, six showed robust smFISH signals in controls that were markedly reduced upon CRISPR knockout (Fig. 4c, e, g, i and Supplementary Fig. 3c, e), and phenotypic analysis was limited to these. For two genes, RNAi based interference studies were performed in the somatic lineage using the c587-GAL4 driver. Phenotypes were quantified by GSC counts, total germ cell numbers, and GSC cell diameter in the apical testis of control and knockout animals (Fig. 4a; Source Data file).

a Schematic of the experimental strategy for TF validation. Lineage-specific GAL4 drivers (carrying UAS-Cas9) were crossed to U6:gRNA lines targeting selected TFs. Crosses were maintained at 18 °C until eclosion, then shifted to 29 °C to induce GAL4-driven expression. b Left: UMAP embedding of the dataset in transcriptional space, with cells colored by lineage-specific latent time. Right: Expression of indicated genes projected onto the UMAP; expression levels are shown as increasing intensity of purple. c Left: Representative smFISH images of Mad transcripts in control and germline-specific Mad knockout (KO) testes. Right: Quantification of Mad smFISH signal across genotypes. d Left: Representative images of control and germline-specific Mad CRISPR mutagenesis testes. Right: Quantification of GSC number, GSC diameter, and total germ cell number. e Left: Representative smFISH images of crc in control and germline-specific crc CRISPR mutagenesis testes. Right: Quantification of crcsmFISH signal. f Left: Control and crc CRISPR mutagenesis testes. Right: Quantification of GSC number, GSC diameter, and total germ cell number. g Left: Representative smFISH images of ovo in control and germline-specific ovo CRISPR mutagenesis testes. Right: Quantification of ovo smFISH signal. h Left: Control and ovo CRISPR mutagenesis testes. Right: Quantification of GSC number, GSC diameter, and total germ cell number. i Left: Representative smFISH images of klu in control and germline-specific klu CRISPR mutagenesis testes. Right: Quantification of klu smFISH signal. j Left: Control and klu CRISPR mutagenesis testes. Right: Quantification of GSC number, GSC diameter, and total germ cell number. In all images, Vasa (green) labels the germline, Fas3 (magenta) marks hub cells, and smFISH signals are shown in white. Scale bars, 20 µm. (c, e, g, i: n = 8), (d, f, h, j: n = 5). Boxplots represent median values within the Q1–Q3 range, while whiskers include the 0–100 percentiles. p values are calculated with two-sided independent t-tests. (*p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001). Exact p values can be found in the Source data file. See also Supplementary Fig. 3, Supplementary Tables 1, 2. Source data is provided as a Source Data file (Fig. 4).
To validate both the testis eRegulons and the CRISPR approach, we first targeted three known stem cell regulators: zfh-1, Stat92E, and Mad5,6,47,59. Knockout of Mad or Stat92E in the germline reduced GSC and total germ cell numbers (Fig. 4d and Supplementary Fig. 3h), consistent with prior reports of BMP and JAK-STAT pathway disruption leading to GSC loss5,6. Although previous Mad mutant studies showed complete germline loss27, the phenotype here was milder, likely due to mosaicism from incomplete CRISPR efficiency. Residual Mad expression in germ cells and preserved expression in the soma (Fig. 4b, c) support this interpretation and confirm lineage specificity of the system. Somatic knockout of zfh1 or Stat92E similarly reduced germ cell numbers (Supplementary Fig. 3d, f), in line with the role of somatic JAK-STAT signaling in GSC support5,47,59. Germline-specific Stat92E knockdown by RNAi testes with an accumulation of CySCs at the hub, recapitulating a previously described phenotype59 (Supplementary Fig. 3g).
Having established the specificity and efficacy of CRISPR mutagenesis, we extended our analysis to additional eRegulon-predicted TFs with unknown testis function (Supplementary Table 1). Knockout of cryptocephal (crc), the Drosophila ortholog of ATF4, in the adult germline reduced GSC and germ cell numbers, and decreased GSC size (Fig. 4f), consistent with its role in sperm production in mice60. Given that crc/ATF4 promotes hematopoietic stem cell maintenance via the integrated stress response (ISR)61, these findings suggest a conserved role for ISR in germline homeostasis. Among the newly tested TFs, ovo and klu emerged as key regulators. Both are expressed in the early germline (Figs. 1g and 4b) and known to function in other stem cell systems41,49,62. While their expression domains partially overlap, ovo is restricted to GSCs and Gbs, whereas klu peaks at the 8 to 16-cell spermatogonial (Sp-8 to 16) stage (Fig. 4b). CRISPR knockout of either gene in adulthood led to striking defects: reduced GSC and germ cell numbers, and smaller GSCs (Fig. 4h, j), highlighting their essential roles in spermatogenesis. We also tested tj, identified by eRegulon analysis and commonly used as a somatic marker, but not further studied functionally in the testis. Adult-specific, soma-targeted tj knockdown, which reduced Tj protein levels in the testis (Supplementary Fig. 3i), significantly reduced GSC and germ cell numbers without strongly affecting GSC size (Supplementary Fig. 3i).
Together, these findings confirm that eRegulon-predicted TFs, including ovo, klu, and crc, are required for adult testis homeostasis and demonstrate the predictive accuracy of our integrated snRNA-seq and snATAC-seq framework.
Multi-modal analysis identifies TF co-regulatory activity
eRegulon predictions, supported by phenotypic similarities from TF perturbation studies, suggested that co-expressed TFs may act through shared regulatory programs. To test this, we quantified overlap in predicted regulatory regions and target genes across eRegulons (see “Methods”). Despite variable overlap, both somatic and germline eRegulons showed substantial convergence (Fig. 5a and Supplementary Fig. 4d), indicating that TFs may coordinate their regulatory function via shared cis-regulatory elements to drive lineage-specific expression programs.

a Heatmap of normalized intersection scores among germline eRegulons based on shared regulatory regions; highlighting overlapping eRegulons in the early germline and the overlap of the achi and vis eRegulons. b ovo and klu eRegulons in GSCs illustrating uniquely and jointly regulated genes and regions. Inset shows the overlap between ovo and klu from (a). c Matrixplot of expression for candidate co-regulated genes. d Pie chart of TF motifs in ovo-klu shared regions; less frequent TFs grouped as Other. Table lists six top enriched motifs. e GO term analysis of uniquely and jointly regulated targets. f stg and nos expression visualized by smFISH in age-matched nos > w1118 (control), nos > kluCRISPR and nos > ovoCRISPR testes with CRISPR interference only performed in adulthood. g Quantification of stg and nos mRNA levels in the respective genetic backgrounds. (g: n = 8 for all experiments). Scale bars = 20 μm. Boxplots represent median values within the Q1–Q3 range, while whiskers include the 1.5*IQR percentiles. p values are calculated with two-sided independent t-tests. (*p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001). Exact p values can be found in the Source data file. See also Supplementary Fig. 4, Supplementary Tables 1, 2 and Supplementary Data 4. Source data is provided as a Source data file (Fig. 5).
A particularly illustrative example of such potential cooperation is provided by the TALE-class homeodomain TFs achintya (achi) and vismay (vis) in secondary spermatocytes (2 °Sc) (Figs. 5a and S5a). These TFs exhibit extensive eRegulon overlap, co-expression, and co-enriched motifs in open chromatin regions, and have been shown to physically interact21. Their interaction with the tMAC complex is essential for regulating spermatocyte-specific gene expression during meiosis and spermatid differentiation21, providing a proof-of-concept that co-expression, motif co-occurrence, and eRegulon intersection can indeed reflect direct combinatorial regulation. GO term enrichment of shared Achi/Vis targets revealed functions in cellular reorganization, cilium motility, and metabolism (Supplementary Fig. 4c), characteristic of other meiotic arrest genes13,19. Motif analysis also identified enrichment for TFIIB in their shared regulatory regions (Supplementary Fig. 4b), consistent with the known promoter-proximal binding of Achi and Vis63.
Encouraged by this, we examined ovo and klu, two TF genes expressed in overlapping early germline populations (Figs. 1g and 4a) and associated with strikingly similar phenotypes upon CRISPR perturbation (Fig. 4h, j). GO term analysis of their shared and unique eRegulon targets implicated both in the regulation of the cell cycle and stem cell differentiation (Fig. 5e). Shared targets included nanos (nos), stg, toutatis (tou), and stem cell tumor (stet) (Fig. 5b), all functionally linked to germline or ovarian stem cell maintenance1,2,3,4,64,65,66. These genes were predominantly expressed in early germline populations (Fig. 5c), consistent with the expression domains of ovo and klu. Genomic regions associated with their shared targets were also enriched for the binding motif of Lola (Fig. 5d), a TF known to be required for GSC maintenance67. To test whether ovo and klu regulate these shared targets, we performed smFISH for nos and stg following germline-specific, adult-only CRISPR knockout of each TF. Both genes were significantly reduced upon adult-stage knockout of ovo or klu in comparison to age-matched control testes (Fig. 5f, g), consistent with their predicted roles in regulating a shared gene module in early germ cells important for GSC maintenance.
Together, these findings suggest that ovo and klu likely participate in a shared transcriptional program critical for early germline function. The interaction between achi and vis illustrates how TF co-expression, motif co-enrichment, and eRegulon overlap can signal physical interaction and functional co-regulation. More broadly, our multimodal single-cell analysis maps of lineage-specific regulatory networks reveal candidate co-regulatory partnerships that may ensure the stability and precision of stem cell gene expression programs.
Ligand-receptor predictions identify Wnt signaling
Numerous signaling pathways regulating stem cell behavior in the Drosophila testis have been well characterized1,2,4,35,68,69,70,71, yet a comprehensive view of cell-cell communication remains incomplete. To address this, we used our high-resolution single-nucleus transcriptome dataset to predict ligand–receptor interactions across testis cell types. We applied the LIANA+ package72 with a curated list of Drosophila ligand-receptor pairs73 to infer signaling networks from gene expression data. This revealed both established and underexplored pathways active in early adult testes, particularly among hub cells, GSCs, and CySCs (Fig. 6a and Supplementary Fig. 5a). For instance, consistent with previous studies7, we detected hh in hub cells and its receptor patched (ptc) in CySCs, confirming somatic Hh–Patched signaling (Fig. 6a). Likewise, gbb expression in hub cells and its receptor tkv in GSCs supports earlier findings that BMP signaling promotes GSC maintenance6. Notably, our analysis also uncovered less-characterized pathways, including Wnt signaling (Fig. 6a and Supplementary Fig. 5a), a central regulator of stem cell function in other tissues such as the ovary74,75,76,77,78,79,80, but not yet systematically studied in the testis81.

a Dotplot of inferred receptor-ligand interactions, grouped by pathway and cell type; dot size and color indicate statistical significance (−log p value) and interaction strength. b Matrixplot of mean expression of Wnt ligands and receptors in GSCs, CySCs, and hub cells. c smFISH validation of Wnt ligand and receptor gene expression in vasaEGFP testes. d Schematic of the lineage-specific RNAi approach to target Wnt pathway components. GAL4 driver lines were crossed to UAS-RNAi lines, with temporal control achieved by temperature shift in adulthood; testes were analyzed after 7 days in age-matched controls and knockdowns. RNAi-mediated knockdown of Wnt ligands or receptors in hub (upd-GAL4;tub-GAL80ts) (e), early germline (nos-GAL4) (g), or soma (c587-GAL4;tub-GAL80ts) (i) affects GSC numbers. f, h, j Quantification of GSC numbers in the respective genotypes. Immunofluorescence markers: Fas3 (hub, magenta), Vasa (germline, green), Tj (soma, yellow). (f: n = 15, 6, 19, 14; h: n = 18, 19, 19, 17; j: n = 17, 15, 16, 19). Scale bars, 20 µm. Boxplots represent median values within the Q1–Q3 range, while whiskers include the 0–100 percentiles. p values are calculated with two-sided independent t-tests. (*p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001). Exact p values can be found in the Source data file. See also Supplementary Fig. 5, Supplementary Table 1, Supplementary Data 4. Source data is provided as a Source data file (Fig. 6).
To validate Wnt pathway predictions from our transcriptomic analysis, we used smFISH to visualize mRNA expression of Wnt components in early testis cell types. Consistent with the snRNA-seq data (Fig. 6b), smFISH confirmed strong DWnt4 and DWnt6 expression in hub cells and CySCs, but little to no signal in germline cells (Fig. 6c). In contrast, components with lower transcript levels in the snRNA-seq data showed variable smFISH signal intensity and spatial distribution. For example, wg and DWnt5 were weakly expressed in hub cells and CySCs by both methods, whereas fz and fz2—the most highly expressed Wnt receptor genes in the niche according to snRNA-seq—exhibited weak but detectable smFISH signals in all niche cells, including GSCs (Fig. 6c). Similarly, the canonical co-receptor gene arrow (arr) and the non-canonical co-receptor gene off-track (otk) were broadly but weakly expressed in all niche cells by snRNA-seq (Fig. 6b), a pattern mirrored by smFISH (Fig. 6c), with otk showing relatively stronger expression in hub cells and CySCs.
To dissect the role of Wnt signaling in adult testis homeostasis, we used gene-specific UAS-RNAi lines driven by lineage-restricted GAL4 drivers (nos-GAL458 for germline, c587-GAL482,83 for the somatic lineage, upd-GAL484 for hub cells) under temporal control (tub-GAL80ts), enabling adult-specific knockdowns following temperature shift (Fig. 6d). Expression and interaction data identified hub cells and CySCs as key sources of Wnt ligands (Fig. 6a–c). Consistent with this, adult-specific DWnt4 depletion in hub cells (upd-GAL4;tub-GAL80ts) significantly reduced GSC numbers, while DWnt6 knockdown increased them (Fig. 6e, f). Strikingly, the reverse was observed when DWnt4 or DWnt6 were depleted in the somatic lineage including CySCs (c587-GAL4;tub-GAL80ts): DWnt6 knockdown reduced GSCs, whereas DWnt4 depletion increased them (Fig. 6i, j), suggesting ligand-specific, lineage-dependent effects on GSC maintenance. To test whether GSCs directly receive Wnt signals, we knocked down fz, fz2, fz4, arr, and otk in the germline using nos-GAL4. Adult-only interference significantly reduced GSC numbers (Fig. 6g, h), indicating that Wnt reception in GSCs is essential for their maintenance. Notably, smFISH revealed low fz, otk, and arr transcript levels in adult GSCs (Fig. 6c), showing that even low expression permits functional Wnt responsiveness and RNAi sensitivity, supported by previous findings85. Mild phenotypes from individual fz knockdowns further suggest receptor redundancy. We next asked whether Wnt reception in somatic cells influences GSC behavior. Knockdown of fz, the most strongly expressed Wnt receptor, in hub cells or the somatic lineage reduced (or increased) GSC numbers (Fig. 6e, f), indicating that Wnt acts on somatic support cells to modulate the niche environment. Similar phenotypes were observed after continuous developmental knockdown (Supplementary Fig. 5b–g) suggest that Wnt signaling primarily contributes to adult GSC homeostasis, though an additional developmental role cannot be fully excluded.
Together, these findings show that somatically derived Wnt ligands from hub cells and CySCs promote GSC maintenance by acting on both germline and soma. Canonical and non-canonical Wnt pathways contribute to this regulation, and even low receptor expression in GSCs ensures responsiveness and supports stem cell maintenance.
Leveraging multimodal data to decode Wnt control in testis
Functional perturbation of Wnt pathway components revealed a critical role for canonical Wnt signaling in GSC maintenance. This pathway requires the co-receptor Arr/LRP6, which stabilizes Armadillo (Arm) to promote its nuclear translocation and convert Pangolin (Pan; TCF/LEF homolog) from a repressor to an activator. A Wnt-responsive reporter driven by multimerized TCF motifs86 was active in GSCs, CySCs, and hub cells (Supplementary Fig. 6a), consistent with Pan eRegulon activity across these populations (Supplementary Fig. 6b, c, g). In GSCs, Pan is predicted to activate how and lola, two genes essential for GSC maintenance67,87 (Supplementary Fig. 6b); in hub cells, it regulates adhesion-related genes including shotgun (shg), Cad87A, and gliolectin (glec)88 (Supplementary Fig. 6c); and in CySCs, it targets genes enriched for signaling functions (Supplementary Fig. 6g).
To assess functional relevance, we performed lineage-specific, adult-only RNAi against pan. Germline knockdown produced two phenotypes with equal frequency: in ~50% of testes, the hub was retained and GSC numbers remained comparable to controls, but total germ cell number was reduced (Fig. 7a–c); in the other half, the hub was lost and single germ cells were markedly increased (Fig. 7d, e). These findings suggest that pan function is required both to sustain germline progenitor expansion and to maintain niche architecture, with the observed variability likely reflecting differences in knockdown efficiency, timing of interference, or niche sensitivity to Wnt pathway perturbation. Notably, these phenotypes were more severe than those seen with hub-derived Wnt ligand depletion (Fig. 6e, f), likely due to (1) redundancy among Wnt ligands (e.g., wg, DWnt4) and (2) the dual role of Pan as a Wnt-dependent activator and repressor89, making its loss more disruptive. Adult-specific knockdown of pan in hub cells did not alter GSC numbers after 7 days (Supplementary Fig. 6e–e”) but increased Fas3-positive (hub) cells by day 10 (Supplementary Fig. 6d, f, f’), indicating that hub cells actively respond to canonical Wnt signaling. This supports our predicted ligand-receptor interactions (Fig. 6a).

a, b Adult-specific knockdown of pan in the germline using nos-GAL4 leads to a mild phenotype in 50% of testes. Immunostaining shows Fas3 (magenta, hub), Vasa (green, germline), and Tj (yellow, soma). c Quantification of GSC number and total germ cell number up to the spermatocyte stage in age-matched control and pan-RNAi testes. d Representative image of a severe phenotype upon adult-only germline-specific pan knockdown, showing hub loss. Immunostaining as in (a). e Quantification of individual germ cells up to the spermatocyte stage under control and experimental conditions. f TF network regulating DWnt6 through 10 regulatory regions in CySCs. TFs are shown in blue, regions in orange, and DWnt6 in red; bold lines highlight Ci binding at region R6 (chr2L:7330491-7330991). g ci eRegulon of in CySCs and hub cells. Putative target regions are shown in orange and their respective target genes in red. Targets shared between CySCs and hub cells (DWnt6, CG15365, CG13506) are circled. R6 is highlighted as a shared regulatory region for wg and DWnt6. h Enriched TF motifs in regulatory region R6. i Chromatin accessibility in GSCs (blue), CySCs (orange), and hub cells (green). Gene bodies of wg and DWnt6 are shown below, with exons in blue and transcriptional orientation indicated by arrows. Inferred regulatory regions are marked in orange. R6 and R7 form a single accessible regulatory region potentially regulating both wg and DWnt6 (green arrows). j dsRed reporter driven by R6 + R7 region shows activity in somatic niche; Fas3 (yellow) marks the hub, Vasa (green) the germline, and mCherry (magenta) the reporter. k smFISH visualization of Wnt6 (white) and wg (green) mRNA in c578>mCherryRNAi and c587>ciRNAi testes. l Quantification of wg and DWnt6 mRNA levels under the indicated genetic conditions. (c: n = 18, 10, 5, 5; e: n = 5 for all; I: n = 5 for all). Scale bars, 20 μm. Boxplots represent median values within the Q1–Q3 range, while whiskers include the 0–100 percentiles. p values are calculated with two-sided independent t-tests. (*p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001) Exact p values can be found in the Source Data file. See also Supplementary Fig. 6, and Supplementary Tables 1, 2, Supplementary Data 4. Source data is provided as a Source Data file (Fig. 7).
Given that Wnt pathway activation in somatic cells is essential for GSC maintenance, we next investigated how the expression of Wnt ligands themselves is regulated within the niche. To this end, we focused on DWnt6, a novel marker of CySCs. eRegulon analysis predicted regulation by nine TFs through 10 distinct regions, including Tj and Dsx (Fig. 7f). The Hh effector Ci was predicted to regulate DWnt6 and wg in CySCs via a shared regulatory element (R6 + R7; Fig. 7g, i), and to regulate DWnt6 in hub cells through the same region, which harbors additional motifs for Br, Tj, Dsx, and CG5953 (Fig. 7h). R6 + R7 is located near the DWnt6 transcription start site and is accessible in CySCs and hub cells, but not GSCs (Fig. 7i). Reporter assays for the R6 + R7 region revealed strong enhancer activity in hub cells and weaker, yet detectable, expression in CySCs, consistent with chromatin accessibility patterns (Fig. 7i, j). Somatic knockdown of ci using c587-GAL4 led to reduced expression of both DWnt6 and wg in hub cells and CySCs (Fig. 7k, l), with DWnt6 showing a more pronounced decrease, suggesting a greater dependence on Ci-mediated input. Together, these findings support a model in which Hh signaling promotes Wnt ligand expression and establishes a regulatory feedback loop between hub and CySCs (Supplementary Fig. 6h).
In summary, our multimodal single-cell analysis reveals a previously unrecognized regulatory circuit linking Hh and Wnt signaling in the Drosophila testis niche. By integrating chromatin accessibility, gene expression, and TF motif analysis, we identify upstream regulators of Wnt ligands and downstream targets of the effector Pan. These data uncover a reciprocal signaling axis between CySCs and hub cells that coordinates stem cell regulation across lineages. More broadly, this work demonstrates how multimodal inference can resolve intercellular signaling logic at single-cell resolution.
link