STInt: An integrated dataset covering science, technology and industry information in the pharmaceutical field
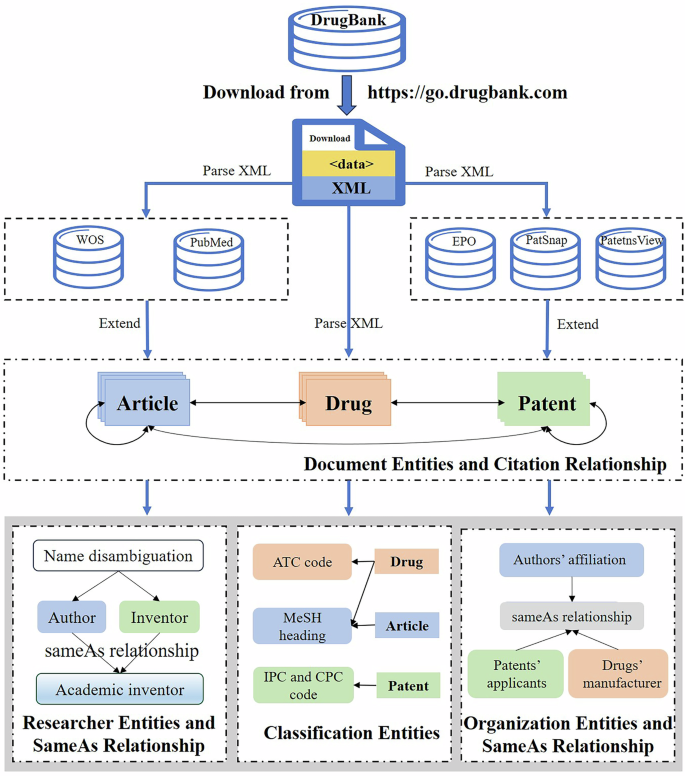
The overall procedure for constructing our STInt dataset is illustrated in Fig. 2. More specifically, the DrugBank database was downloaded in the format of XML from its website ( on 1 November 2019 at the first place. After parsing the XML file, one can readily obtain the drug-related information (such as name, type, description, and the resulting synonyms), and the PMID and patent numbers of associated academic articles and patents. Further, the article and patent document entities are expanded from multiple databases (such as PubMed, Web of Science, and EPO) based on PMID and patent numbers. In the meanwhile, the citation relationships among the article, patent, and drug entities are constructed. Finally, the researcher and organization entities are extracted and disambiguated from fetched articles and patents, apart from MeSH headings, WoS categories, ATC, IPC and CPC codes. In addition, after disambiguation, the sameAs relations between researcher entities and those between organization entities are identified. In the following subsections, several core modules in Fig. 2 will be described in more detail.
Multi-source integrated dataset construction process.
Document entities and citation relations
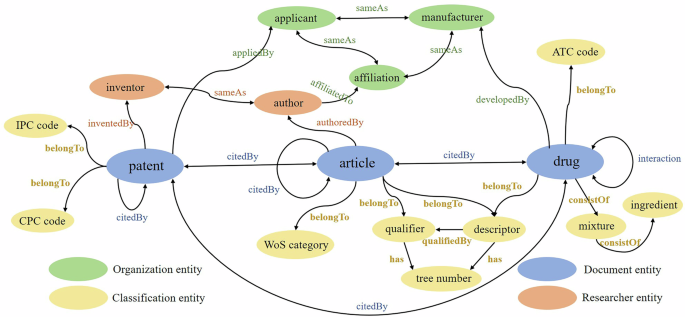
In this paper, we extract three types of document entities (drugs, articles, and patents) from downloaded XML files, and further, eight citation relationships among three types of entities are constructed.
A snippet of XML file is shown in Fig. 3, which describes the related information (such as drugbank id, description, and synonyms) of the drug Lepirudin, as well as information on articles and patents cited by this drug. Note that all drugs are further categorized into biotech and small molecule in our dataset. Then, the related information about academic articles and patents (such as title, abstract, and references) is fetched from PubMed, WoS, EPO, PatSnap, and PatentsView databases by following the procedure in Fig. 4.
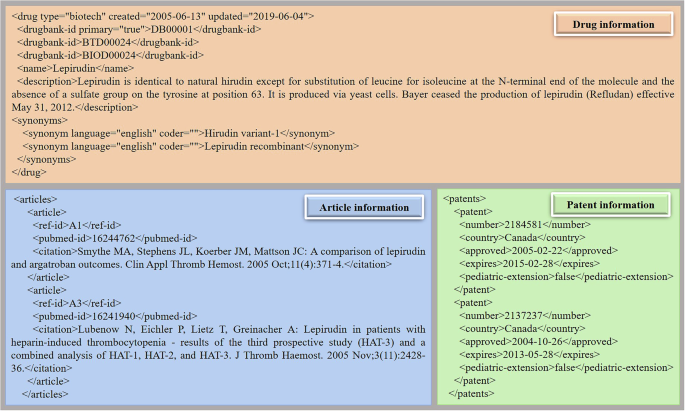

A snippet of XML file related to drug information, and its associated articles and patents.
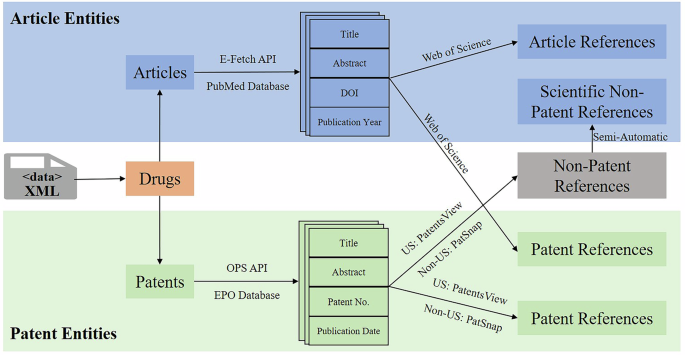
Procedure of extending the information of article and patent entities.
Article entities
The academic articles in our STInt dataset have three groups as follows: (a) academic articles cited by drugs, (b) references of academic articles cited by drugs, and (c) scientific non-patent references (sNPRs) of patents cited by drugs.
First, the PMID of each article cited by a drug is extracted from downloaded XML file. According to the PMID, the relevant article information, including title, abstract, author, DOI, and publication year, is obtained from the PubMed literature database through the E-Fetch API (https://www.ncbi.nlm.nih.gov/books/NBK25499/#chapter4.EFetch).
Second, after the reference list of each article cited by drugs is obtained through the WoS database according to the resulting PMID or DOI, the DOI of each reference is cleaned by the approach in Xu et al.30. Then, the related information (e.g., title, abstract, publication year, and so on) of each reference is further fetched from the WoS and PubMed databases according to the cleaned DOI.
As for sNPRs of patents cited by drugs, the resulting DOI or PMID of each sNPR is obtained by manually searching in the first place. Before this, we filter the following references according to manually curated rules, such as FDA guidelines, product labels, lab manuals, etc. Finally, the title, abstract, publication year and the other information of each sNPR can be readily obtained through the WoS and PubMed databases. For more elaborate and detailed descriptions, we refer the readers to Xu et al.28.
Patent entities
Similar to the article entities, the patent entities in our STInt dataset have also three groups: (a) patents cited by drugs, (b) patents that are cited by patents cited by drugs, and (c) patents that are cited by articles cited by drugs.
First, this paper extracts patent number of each patent cited by a drug from downloaded XML file. According to the patent number, the relevant patent information, including the title, abstract, inventor, applicant, and publication date, is fetched from the EPO patent database through the OPS API (http://ops.epo.org).
Second, to obtain the related information of the patents that are cited by patents cited by drugs, two patent databases, PatentsView and PatSnap, are involved. More specifically, the U.S. patent information comes from the PatentsView database, and the non-U.S. patent information from the PatSnap database.
Finally, a small number of cited patents usually appear in the reference lists of the articles cited by drugs. The resulting patent numbers of the cited patents are extracted according to manually curated rules. Then, the information about these patents is obtained from the PatSnap database. Before this, the patent numbers are manually reviewed one by one.
Constructing citation relations
The citation relations in our dataset can be divided into eight types of relationships: (1) citations between articles, (2) citations from articles to patents, (3) citations from articles to drugs, (4) citations between patents, (5) citations from patents to articles, (6) citations from patents to drugs, (7) citations from drugs to articles, (8) citations from drugs to patents. The process of constructing citation relations is shown in Fig. 5.
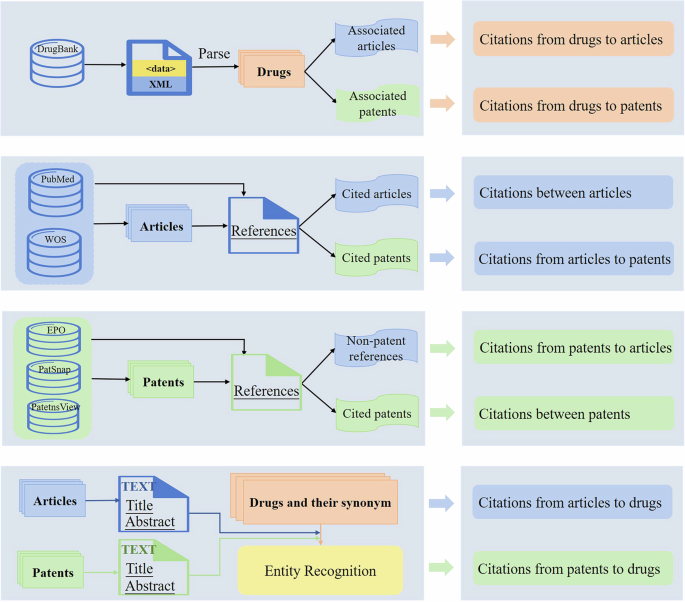
The process of constructing citation data.
This paper constructs the citations from drugs to articles and those from drugs to patents based on the articles and patents information associated with drugs in the downloaded XML file. According to the reference list of each article cited by drugs, the citations between articles and those from articles to patents are constructed. Similarly, according to the reference list of each patent cited by drugs, the citations between patents and those from patents to articles are constructed.
In general, no drug appears in the reference list of an article or a patent. To construct the citations from articles to drugs and those from patents to drugs, we use drug entity recognition methods to extract entity mentions from the corresponding title and abstract of each article/patent cited by drugs. If an article/patent mentions a drug or its synonyms, a citation relation from this article/patent to this drug is constructed. For specific detail about drug entity recognition, please refer to Xu et al.8,22.
Researcher disambiguation and sameAs relationship
It is well known that many different researchers share the same name (viz., the homonym problem), and individual researchers sometimes publish their works (such as academic articles and patents) under different names (i.e., the synonym problem). This is known as name ambiguity problem in the literature31. A revised rule-based scoring and clustering method32 is utilized in this study for disambiguating the authors. As for the inventors, we utilize a semi-automatic method to split and check the first and last names of each inventor, and then the inventors are disambiguated by several manually curated rules on the basis of the applicants, co-inventors, address, theme of the resulting patent, and so on.
Further, we construct the sameAs relationship between authors and inventors, i.e., academic inventors15,33,34. The academic inventors are known to author scientific publications and patent inventions simultaneously, and they usually play the role of gatekeepers between science and technology. The procedure of name disambiguation and sameAs relationship construction is shown in Fig. 6. For more detail on name disambiguation and identification of academic inventors, we refer the readers to Xu et al.16.
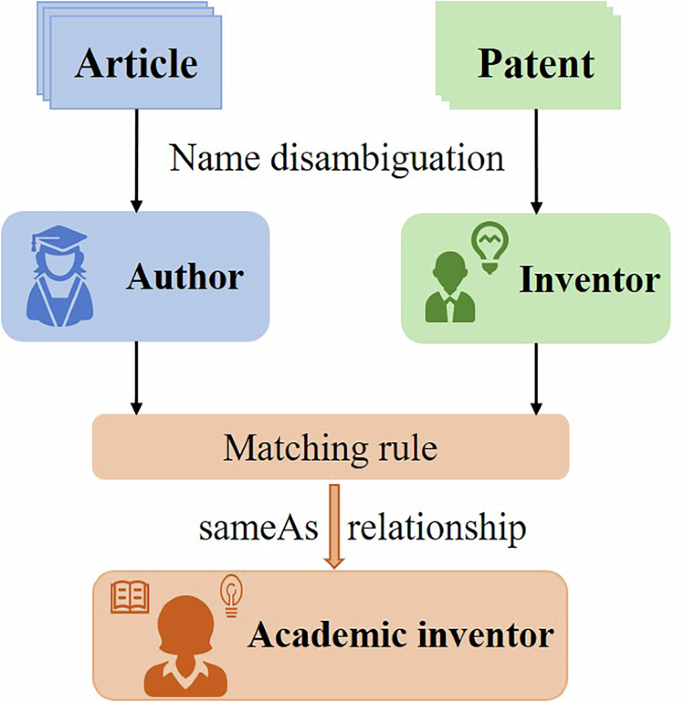
The process of researcher disambiguation and sameAs relationship identification.
Organization disambiguation and sameAs relationship
The organization entities in our STInt dataset include the affiliations in scientific publications, the applicants in patents, and the manufactures of drugs. The affiliations are extracted from the byline information of each article, the applicants from each patent document, and the manufactures from the downloaded XML file. In addition, we construct the sameAs relationship among these three types of organizations.
Similar to researcher entities, an organization may be expressed differently in the literature. Therefore, this study disambiguates three types of organizations separately. More specifically, the similarity based on Levenshtein distance35 is calculated to identify the organizations with similar names. The following steps are involved in the procedure of similarity calculation. (1) Tokenization: the organization names are tokenized into multiple tokens. (2) Sorting: the segmented tokens are sorted. (3) Merging: the sorted tokens are recombined into a string. (4) Levenshtein distance calculation, also known as the edit distance, which can measure the difference between two strings. In principle, this distance represents the minimum number of editing operations (insertion, deletion, and substitution) required to convert one string into another. (5) Similarity score: The similarity score is calculated on the basis of Levenshtein distance, which ranges from 0 to 100, with a higher score indicating a higher degree of similarity. The similarity score can be formally defined as follows.
$${{Score}}_{{similarity}}\left({o}_{1},{o}_{2}\right)=\left(1-\frac{{distance}\left({o}_{1},{o}_{2}\right)}{{Max}(\left|{o}_{1}\right|,\left|{o}_{2}\right|)}\right)\times 100$$
Here, \(\left|{o}_{1}\right|\) and \(\left|{o}_{2}\right|\) denote the number of characters in the names of organizations \({o}_{1}\) and \({o}_{2}\) respectively, and \({distance}\left({o}_{1},{o}_{2}\right)\) represents the Levenshtein distance between the names of organizations \({o}_{1}\) and \({o}_{2}\).
By trial and error, the similarity threshold is fixed to 90 in this work. That is, each pair of organizations with similarity greater than 90 is manually checked on whether or not the same organization is involved. For ease of understanding, several examples are shown in Table 1. From Table 1, it is easy to find that in the pairs 1, 2 and 5, the same organization is involved, but in the pairs 3, 4 and 6, different organizations are observed. If the same organization is observed, the resulting names will be combined. For instance, the organizations with ID = 1 and 8279 are merged into “Utrecht University”.
After disambiguating all organizations, we find that several organizations simultaneously participate in two/three activities of scientific research, technological development, and industrial progress. These organizations usually play a crucial role in the interactions among science, technology, and industry. Therefore, we further identify such organizations, and the sameAs relationship is formed between the resulting organizations. It is noteworthy that an organization may be expressed inconsistently across three types of organizations, for example, Novartis Pharmaceuticals Corporation in the affiliations versus Novartis pharmaceuticals corp in the manufacturers. Again, similarity calculation method with a different threshold 85 is utilized in this study.
Classification entities
The classification entities in our dataset include ATC codes for drugs, MeSH headings for articles and drugs, WoS categories for articles, and IPC and CPC codes for patents. Just as its name implies, WoS category comes from the WoS database. No complicated structure is involved in the WoS category, so the construction process of the other classification entities will be described below.
ATC code for drugs
The ATC classification system is an international standard maintained by the World Health Organization (WHO) for coding each drug. It standardizes the classification of drugs through a five-level classification system (including anatomical main groups, therapeutic/pharmacological subgroups, therapeutic subgroups, chemical/therapeutic/pharmacological subgroups, and specific chemical substances), thus allowing for the uniform reference of drugs in different countries/regions. The ATC system is not only widely used in drug regulation, drug policy development, pharmacy research, and education, but it also facilitates the communication of drug use and policy at the international level by providing a common language. The importance of the ATC classification system lies in the fact that it provides accuracy and consistency of drug information on a global scale, thereby improving the quality of drug use and management, supporting the safe and effective use of drugs, and facilitating the international exchange of information on drug use.
Fig. 7 shows a snippet of ATC code of the drug (Lepirudin) in the downloaded XML file. It is not difficult to see that this XML fragment describes the ATC code of a focal drug and its corresponding level. In the XML fragment, the element “ < atc-code > ” has an attribute code = “B01AE02”, which is the ATC code of a particular drug. In addition, within the “ < atc-code > ” element, there are multiple “ < level > ” elements, each representing a different level in the ATC classification system, which progressively expand from more specific classifications to broader ones, as shown in Fig. 7 for the ATC code as an example:
-
code = “B01AE”: indicates direct thrombin inhibitors.
-
code = “B01A” and code = “B01”: both denote ANTITHROMBOTIC AGENTS, where the descriptions are the same for both tiers, as this particular subcategory is not further subdivided in the ATC system.
-
code = “B”: indicates BLOOD AND BLOOD FORMING ORGANS, which is the first level of classification in the ATC classification system.

A snippet of ATC code of the drug Lepirudin in the downloaded XML file.
Thus, “B01AE02” in Fig. 7 is classified as a direct thrombin inhibitor, which belongs to the class of antithrombotic agents and is ultimately classified as a blood and hematopoietic organ related drug.
MeSH headings for articles and drugs
The MeSH is a standardized medical terminology system maintained by the U.S. National Library of Medicine (NLM), which is mainly used for indexing, cataloging, and retrieval of literature in medicine, biomedicine, public health, and related fields. The MeSH provides a unified set of terms to describe medical topics and concepts, which mainly includes descriptors (core concepts), qualifiers (aspects of refinement), and index terms (synonyms or related terms) to resolve confusion caused by different terms and synonyms. It is widely used for indexing literature in medical libraries and databases such as PubMed. The importance of this system is to provide a uniform, standardized language for the medical and biomedical fields, thus facilitating the effective sharing and application of medical knowledge.
In the MeSH system, the descriptor and qualifier are elements used to describe key topics or terms in the medical or biological literature that have specific meanings and uses in the MeSH system. The descriptors represent the key concepts or topics discussed in the literature. These descriptors are predefined terms in the MeSH system, which are used to categorize and index the theme of the medical literature. The qualifiers are used to further qualify the descriptors, providing more specific detail about the resulting descriptors, such as a particular aspect, attribute, or a particular perspective of the descriptor discussed in the literature. The difference between them is that a descriptor defines the main discussion points of a scholarly article, while a qualifier provides more specific contextual or aspect-specific information about those discussion points. In the MeSH system, the descriptors and qualifiers are used together to more precisely define the content and focus of the literature.
Fig. 8 illustrates the MeSH heading information for the article with DOI = 10.1007/BF00421005 and drug Lepirudin. This XML snippet describes the medical subject terms of the article, containing the DescriptorName and QualifierName. In Fig. 8, Adult, Anti-Anxiety Agents, Azepines, and Thiophenes are all descriptors that identify the main theme. The qualifier associated with Anti-Anxiety Agents is pharmacology, which indicates that the discussion of anxiolytic drugs is from a pharmacological perspective. At the same time, each term has a unique identifier (UI) and MajorTopicYN indicates whether or not it is the main topic. Notably, only descriptor information is involved for each drug (c.f. Fig. 8). Our STInt dataset covers all these information.
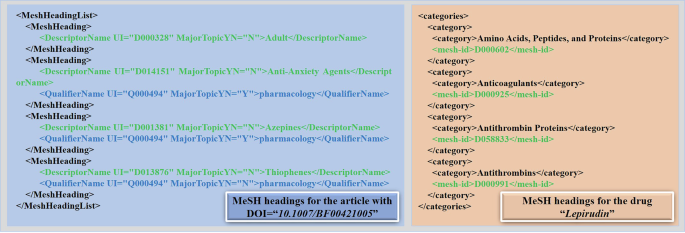

A snippet of XML file relevant to MeSH headings of an article and a drug.
It is well known that the MeSH descriptors are organized in 16 categories: category A for anatomic terms, category B for organisms, and so on. Each category is further divided into sub-categories. Within each sub-category, the descriptors are arranged hierarchically from most general to most specific in up to thirteen hierarchical levels. However, the UI in Fig. 8 cannot reflect any hierarchical structure information. For this purpose, we download the complete MeSH data from In addition to parsing the descriptors and qualifiers from the downloaded MeSH file, we also parse the resulting tree number for each descriptor and qualifier in our STInt dataset. In the MeSH system, tree numbers are key organizational and categorization tools used to specify the position of descriptors and qualifiers in the medical terminology hierarchy. These tree numbers not only reflect the categorization of different medical and biomedical fields, but also reveal the relationships between terms. The descriptor tree numbers can guide the user through the location of a particular descriptor within the overall medical subject system and its association with other descriptors, while the qualifier tree numbers help to identify the classification of qualifiers and how they are associated with descriptors.
IPC and CPC codes for patents
The IPC is an international patent classification system managed by the World Intellectual Property Organization (WIPO) for classifying patents and utility models according to technical fields. Its structure consists of multiple levels: section, class, subclass, main group, and subgroup, allowing for the subdivision from broad fields of technology to specific technological realizations. The CPC is a detailed patent classification jointly sponsored by the European Patent Office (EPO) and the United States Patent and Trademark Office (USPTO). The structure of the CPC draws on the IPC and introduces additional levels of subdivision and labels for classification, including section, class, subclass, main group, subgroup, and other more specific classification levels.
Fig. 9 shows a snippet of XML file related to the IPC and CPC information of the patent with No. CA1323836. From Fig. 9, three IPC codes are extracted herein: A61K31/22, A61K9/20, and A61K47/02. Each IPC code denotes a specific technical field to which the patented invention belongs. In this example, A61K31/22, A61K9/20 and A61K47/02 denote the field of pharmaceuticals, preparation of pharmaceuticals, and cosmetics or skin care products respectively. In the meanwhile, three CPC codes can also be extracted from the XML snippet in Fig. 9 as follows: A61K9/2004, A61K9/2009, and A61K31/19, which indicate that the patented invention belongs to the field of drug preparation methods or medical devices.
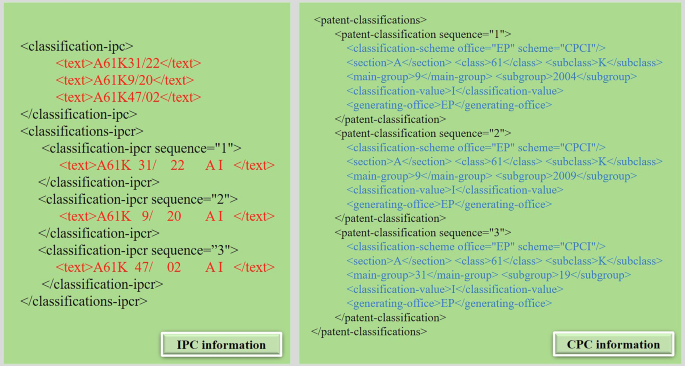

A snippet of XML file related to the IPC and CPC classification information.
link