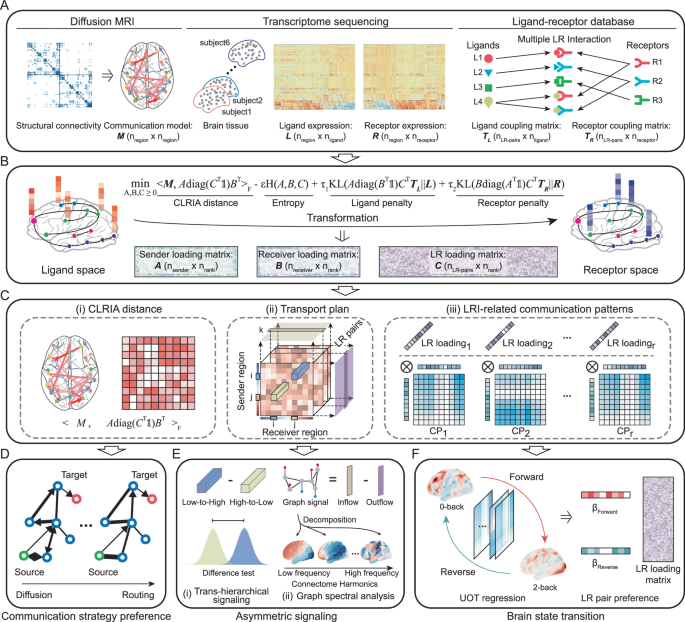
The full details of the theoretical background and implementation can be found in the Supplementary Information.
Ethical statement
We used the publicly available data that participated in the Human Connectome Project (HCP; S1200), and had complete MRI data including diffusion-weighted MRI, all four sessions of resting-state fMRI and task fMRI related to working memory. All participants provided informed consent as part of the HCP, and the study protocol was approved by the institutional review boards of the WU-Minn HCP Consortium. The processing of these data was approved by the medical ethical committee of the Charité Medical Center in Berlin. Ethics oversight of gene expression data was performed by the Allen institute for brain science.
HCP dataset
Structural connectivity
SC was inferred from minimally preprocessed, high-resolution diffusion-weighted MRI data from healthy young adults participating in the Human Connectome Project. Details of the diffusion MRI acquisition and preprocessing are described in Sotiropoulos et al.83 and Glasser et al.84 After estimating the orientation distribution function using a multi-shell, multi-tissue constrained spherical deconvolution model85, whole-brain white matter tractograms of individual participants were mapped using MRtrix386 with the following parameters: anatomically constrained tractography (ACT87), iFOD2 tracking algorithm88, 5M streamlines with dynamic white-matter seeding, 300 mm maximum streamline length, and 0.1 FOD amplitude cutoff for termination. To better match the whole-brain tractograms to the diffusion properties of the observed data, we also computed streamline weights, which are designed to reduce known biases in tractography data (SIFT289). The human Brainnetome atlas90 divides the cortex and subcortex into 246 regions. Connection strength between any two regions was the SIFT2-weighted sum of the streamlines connecting those regions divided by the total gray matter volume of those regions. The strengths with fewer than 10 streamlines were pruned to reduce the high false positive rate. The group-representative connectivity matrix was computed using a consensus method that combined individual-level networks while preserving the average connection density across subjects (N = 1061, 487 males, 574 females, age 22–36)91.
Functional connectivity
FC was mapped using minimally preprocessed, ICA-FIX resting-state functional MRI data from the same sample of HCP participants. For each subject, four 14 min and 33 s scans (0.72 s TR) were acquired following details described in Glasser et al.84 and Smith et al.92 Following another study93, motion and global signal outlier timepoints were identified using an approach adapted from Artifact Detection Tools (ART) from the CONN Toolbox94. After removing global signaling outliers and the first 10 volumes from each scan, additional nuisance variables, e.g., including an offset term, linear trend, 6 motion parameters and their derivatives, squares, and squared derivatives (24 motion regressors), and 10 anatomical CompCor (aCompCor) regressors, were regressed out. FC matrices were calculated using the Pearson correlation between each region-pair’s average time series in the Brainnetome atlas90, resulting in 4 FC matrices for each subject. A final group-representative FC matrix was computed as the average within subjects and then across subjects (N = 1096, 500 males, 596 females, age 22–36).
Contrast map of task fMRI
The task-evoked responses were already processed by minimal preprocessing pipelines84. Here, we mainly focused on general linear model contrasts from working memory (0 back and 2 back). We parcellated these maps into 246 regions using the Brainnetome atlas90 for each subject (N = 1080, 494 males, 586 females, age 22–36) and extracted the average value from each region-of-interests (ROI). Because we mainly focused on the active region, the regions with negative average values were set as zero.
AHBA gene expression processing
Human brain gene expression from AHBA ( was measured in six post-mortem brains, containing 3702 spatially distinct samples and 58692 probes95. Of note, the original MNI coordinates did not take into account nonlinear deformation, so we used the coordinates from Devenyi ( which were registered to MNI space (ICBM 2009c). Further processing was guided by Arnatkevic et al.96 and abagen97. First, probes with intensities below the background in at least 50% of the samples were excluded. Based on the updated probe-to-gene annotations from Re-annotator98, probe selection was performed based on differential stability99. Next, expression normalization was based on the robust sigmoid method, which is robust to outliers. Finally, samples assigned to the human Brainnetome atlas90 within 2 mm Euclidian distance of a parcel were averaged first within each donor and then across the donors. Because only two of the six donors included samples from the right hemisphere, only samples from the left hemisphere were considered100, resulting in an expression matrix of 123 regions and 15634 genes.
Ligand receptor pairs
CellChatDB v245 is an updated version of the original CellChatDB13, which was expanded by more than 1000 protein and non-protein interactions based on peer reviewed literature and existing databases, such as CellPhoneDB12 and NeuronChatDB21 (metabolic and synaptic signaling). This updated version contains a total of 3237 unique pairs, 1102 of which are neurotransmitters. To account for the communication between brain regions, we only retained secreted and non-protein (mainly synaptic signaling) pairs. Furthermore, ligand-receptor pairs (LR-pairs) not detected in the AHBA dataset were excluded, resulting in a final list of 1242 pairs, 663 of which are neurotransmitters. The geometric mean of their related genes was used to represent the amount of ligand or receptor.
Brainspan developmental gene expression processing
The Brainspan database quantifies the gene expression in the brain throughout development ( and includes postmortem brains from 8 post-conception weeks to 40 years58. RNA-seq data were used to explore the expression changes along development. Referring to Hansen, et al.100, we manually binned the ages into six periods: fetus (8–37 post-conception weeks), infant (4 months–1 year), child (2–8 years), adolescent (11–19 years), young adult (21–30 years), and adult (36–40 years). In each brain region and developmental stage, the log-transformed RPKMs (reads per kilobase per million mapped reads), which measure the expression of genes, were averaged across all the samples.
CLRIA
Interregional communication is constrained by the internal complex connectivity, which make signaling efficient. Ligands and receptors are major signaling molecules during the communication process. If we regarded the expression of ligands and receptors as two distributions over the brain region, then the process of BNC can be viewed as a distributional transformation from ligand to receptor constrained by the underlying anatomical connectivity. From this point of view, we initiated the following multiple sources, multiple targets, optimal transport problem to construct brain interregional communication networks through predefined LR-pairs:
$$I\right_F\left(\right):=\left_^I\right{\left\langle M,\left_\left\right\rangle }_$$
where \(M\in \left_N\times N\times \left^I\right\) is the transport cost constructed from the BNCM; \({{I\right}}_\in I\right_^\left\) is the transport plan of ith LR-pair, indicating the signaling strength between brain regions; \(\left|I\right|\) is the total number of LR-pairs; \(_N\times N\times \left\) is the Frobenius inner product for the matrix; \(^\in {\left}^\) and \(I\right^I\right\in {I}^I\right\) are the amount of ligand that interacted with the cognate receptor and vice versa; and \(\Pi (L^\left,R^ )=\left\{\,\left.{{{\mathcal }}}\in {{\mathbb}}_{+}^{N\times N\times \left|I\right|}\right|{{{\mathcal{P}}}}\times 2{{\mathbb{1}}}_{N}=L{\prime},{{{\mathcal{P}}}}\times 1{{\mathbb{1}}}_{N}=R{\prime} \right\}\) is the mass preserving constraint set.
Of note, current transcriptome sequencing cannot quantify the proportions of their interactions, so we used the total expression in each brain region as a surrogate. Since the total expression and interaction part of a ligand/receptor pair may be different, we relaxed the marginal constraints by adding a KL divergence penalty term on an objective function following Chizat et al.41 We further introduced two coupling matrices \({T}_{L},{T}_{R}\), which were derived from the ligand receptor database (see Supplementary material for further details). Finally, we introduced nonnegative tensor decomposition on the transport plan as a low rank constraint for both computational efficiency and biological significance. Intuitively, this constraint reduces the number of variables to be optimized, which naturally facilitates the computational efficiency. Moreover, a recent solver of the single source-target optimal transport problem benefitted from the low rank properties of the transport cost and transplant plan and achieved remarkable success with respect to efficiency and scalability31,33,34,35. In addition, internal associations between LR-pairs, such as multiple species interactions19 and similar gene compositions, essentially make the transport plan into a low-rank tensor. In addition, tensor decomposition (e.g., tensor component analysis, TCA) exhibits superior performance in latent pattern recovery for multi-dimensional data36,37,38,39. Therefore, we expected that this low-rank constraint would allow us to gain details about the underlying communication patterns. Considering all the above, the final optimal transport problem can be reformulated as:
$${\min }_{\left(A,B,C\right)\in \Pi \left(r\right)}F\left(A,B,C\right):= {\left\langle M,A{{\mbox{diag}}}\left({C}^{T}{{\mathbb{1}}}_{\left|I\right|}\right){B}^{T}\right\rangle }_{F}-\varepsilon {\mbox {H}}\left(A,B,C\right) \\ +{\tau }_{1}{{\mbox{KL}}}\left(A{{\mbox{diag}}}\left({B}^{T}{{\mathbb{1}}}_{N}\right){C}^{T}{T}_{L}\parallel L\right) \\ +{\tau }_{2}{{\mbox{KL}}}\left(B{{\mbox{diag}}}\left({A}^{T}{{\mathbb{1}}}_{N}\right){C}^{T}{T}_{R}\parallel R\right)$$
where A, B and C are factor matrices decomposed from a transport plan \({{{\mathcal{P}}}}\); \({\mbox {H}}\left(A,B,C\right)={\mbox {H}}\left(A\right)+{\mbox {H}}\left(B\right)+{\mbox {H}}(C)\) is the entropy function; and \(\varepsilon,{\tau }_{1},{\tau }_{2}\) are coefficients of regularization terms. \(\Pi \left(r\right)=\left\{\left(A,B,C\right)\in {{\mathbb{R}}}_{+}^{N\times r}\times {{\mathbb{R}}}_{+}^{N\times r}\times {{\mathbb{R}}}_{+}^{\left|I\right|\times r}\right\}\) is the nonnegative constraint on the factor matrix. \(L\in {{\mathbb{R}}}^{N\times m}\) and \(R\in {{\mathbb{R}}}^{N\times n}\) are the ligand and receptor expression matrices in each brain region, respectively. For heteromeric ligands or receptors, the geometric mean of the composed genes was adopted to represent the abundance of ligands or receptors7,101,102. We manually replaced zero with a small number (10−80 as default) in the expression matrix to avoid infinity in the objective function.
BMM algorithm for CLRIA
Of note, KL terms in the objective are not well defined at \({\left(A{{\mbox{diag}}}\left({B}^{T}{{\mathbb{1}}}_{N}\right){C}^{T}{T}_{L}\right)}_{{ij}}=0\) or \({\left(B{{\mbox{diag}}}\left({A}^{T}{{\mathbb{1}}}_{N}\right){C}^{T}{T}_{R}\right)}_{{ij}}=0\), which makes the convergence analysis challenging103. Therefore, we focused on the solution in a perturbed constraint set \({\Pi }_{{{{\rm{\delta }}}}}\left(r\right)=\left\{\left(A,B,C\right)\in {{\mathbb{R}}}_{\delta+}^{N\times r}\times {{\mathbb{R}}}_{\delta+}^{N\times r}\times {{\mathbb{R}}}_{\delta+}^{N\times \left|I\right|}\right\}\), with \(\delta={10}^{-100}\) as the default. Considering the block convexity of \(F\left(A,B,C\right)\) and the interaction term existing in KL divergence, we used a BMM scheme to solve the perturbed problem. The first step was to construct the surrogate function for each block. This step separated the variables and retrieved a closed form solution. Following [Lemma D 4] in the Supplemental material, we can get the surrogate function for the KL term. Based on their properties ([Proposition D 1] in the Supplemental material), we can directly get the surrogate function for each block. Starting from random initialization \(\left({A}^{(0)},\,{B}^{\left(0\right)},\,{C}^{\left(0\right)}\right)\in {\Pi }_{{{{\rm{\delta }}}}}\left(r\right)\), we have the following updates by minimizing each subproblem for \(k\ge 0\):
$$\begin{array}{c}{C}^{\left(k+1\right)}=\max \left\{{C}^{\left(k\right)}\odot \exp \left(-{\Omega }_{C}/{\Lambda }_{C}\right),\delta \right\}\\ {A}^{\left(k+1\right)}=\max \left\{{A}^{\left(k\right)}\odot \exp \left(-{\Omega }_{A}/{\Lambda }_{A}\right),\delta \right\}\\ {B}^{\left(k+1\right)}=\max \left\{{B}^{\left(k\right)}\odot \exp \left(-{\Omega }_{B}/{\Lambda }_{B}\right),\delta \right\}\end{array}$$
where
$${H}_{1}= A{{\mbox{diag}}}\left({B}^{T}{{\mathbb{1}}}_{N}\right){C}^{T}{T}_{L};{H}_{2}=B{{\mbox{diag}}}\left({A}^{T}{{\mathbb{1}}}_{N}\right){C}^{T}{T}_{R}\\ {\Lambda }_{C}= {\tau }_{1}{T}_{L}{{\mathbb{1}}}_{m\times N}A{{\mbox{diag}}}\left({B}^{T}{{\mathbb{1}}}_{N}\right)+{\tau }_{2}{T}_{R}{{\mathbb{1}}}_{n\times N}B{{\mbox{diag}}}\left({A}^{T}{{\mathbb{1}}}_{N}\right)\\ {\Omega }_{C}= {{\mathbb{1}}}_{\left|I\right|}{{\mathscr{D}}}\left({A}^{T}{MB}\right)+{\tau }_{1}{T}_{L}\log {\left(\frac{{H}_{1}}{L}\right)}^{T}A{{\mbox{diag}}}\left({B}^{T}{{\mathbb{1}}}_{N}\right)+{\tau }_{1}{T}_{R}\log {\left(\frac{{H}_{2}}{R}\right)}^{T}B{{\mbox{diag}}}\left({A}^{T}{{\mathbb{1}}}_{N}\right)\\ {\Lambda }_{A}= {\tau }_{1}{{\mathbb{1}}}_{N\times m}{T}_{L}^{T}C{{\mbox{diag}}}\left({B}^{T}{{\mathbb{1}}}_{N}\right)+{\tau }_{2}{{\mathbb{1}}}_{N}{\left({B}^{T}{{\mathbb{1}}}_{N}\odot {C}^{T}{T}_{R}{{\mathbb{1}}}_{n}\right)}^{T}\\ {\Lambda }_{A}= {MB}{{\mbox{diag}}}\left({C}^{T}{{\mathbb{1}}}_{\left|I\right|}\right)+{\tau }_{1}\log \left(\frac{{H}_{1}}{L}\right){T}_{L}^{T}C{{\mbox{diag}}}\left({B}^{T}{{\mathbb{1}}}_{N}\right)+{\tau }_{2}{{\mathbb{1}}}_{N}{{\mathscr{D}}}\left({C}^{T}{T}_{R}\log {\left(\frac{{H}_{2}}{R}\right)}^{T}B\right)\\ {\Lambda }_{B}= {\tau }_{1}{{\mathbb{1}}}_{N}{\left({A}^{T}{{\mathbb{1}}}_{N}\odot {C}^{T}{T}_{L}{{\mathbb{1}}}_{m}\right)}^{T}+{\tau }_{2}{{\mathbb{1}}}_{N\times n}{T}_{R}^{T}C{{\mbox{diag}}}\left({A}^{T}{{\mathbb{1}}}_{N}\right)\\ {\Omega }_{B}= {M}^{T}A{{\mbox{diag}}}\left({C}^{T}{{\mathbb{1}}}_{\left|I\right|}\right)+{\tau }_{1}{{\mathbb{1}}}_{N}{{\mathscr{D}}}\left({C}^{T}{T}_{L}\log {\left(\frac{{H}_{1}}{L}\right)}^{T}A\right)+{\tau }_{2}\log \left(\frac{{H}_{2}}{R}\right){T}_{R}^{T}C{{\mbox{diag}}}\left({A}^{T}{{\mathbb{1}}}_{N}\right)$$
The superscript for each iteration was ignored for simplicity. The derivation of the algorithm is described in the Supplemental material [Lemma 2]. The global convergence of the update rules can be proved using Zangwill’s theorem42 ([Theorem 1] in the Supplemental material).
Computational cost
Given a specific species (e.g., human), the number of LR-pairs \({|I|}\) and related genes of LR-pairs \(m,{n}\) can be regarded as constants. Each iteration was mainly involved with vector/matrix multiplication, which requires \({{{\mathscr{O}}}}\left({N}^{2}r+N\left({mr}+n\right)\right)\), \({{{\mathscr{O}}}}\left({N}^{2}r+N\left({nr}+m\right)\right)\),\({{{\rm{and}}}}\) \({{{\mathscr{O}}}}\left({N}^{2}r+{Nr}\left(m+n\right)\right)\) operations to update \(A,B,C\). Writing K as the number of iterations of the BMM scheme, we ended up with a total computational cost of \({{{\mathscr{O}}}}\left(K{N}^{2}r+KN\left({mr}+{nr}\right)\right)\). If we further use factor matrices \({M}_{1}\in {{\mathbb{R}}}^{N\times d}\) and \({M}_{2}\in {{\mathbb{R}}}^{N\times d}\) such that \(M={M}_{1}{M}_{2}^{T}\), which is usually meaningful for brain networks64,104, then the algorithm requires \({{{\mathscr{O}}}}\left(KN\left({dr}+{mr}+{nr}\right)\right)\) operations. This linear cost with respect to the number of brain regions would facilitate the exploration of brain communication on a finer scale.
Comparison with other methods
We compared CLRIA against several established cell communication frameworks, including CellPhone12, CellChat13,45, GeoMean46, CellLinker20, MEBOCOST22 and NeuronChat21. CellPhone and CellChat are popular software packages for cell-cell communication analysis using single cell sequencing data, and their recent versions support distance constraints. GeoMean is short for geometric mean, which is commonly used to calculate a communication score for each LR pair between different cell types. The incorporation of metabolic processes associated with the synthesis of non-protein ligands constitutes a pivotal feature of CellLinker, MEBOCOST and NeuronChat. The ligand-receptor database ultized by CellPhone, CellChat and geometric mean was identical as CLRIA. In contrast, the self-curated ligand-receptor database were used for CellLinker, MEBOCOST and NeuronChat. To ensure compatibility with the input requirements of these benchmarked methods, brain regions were computationally processed as different cell types. Following the acquisition of the communication score, it is possible to infer the direction of signaling flow between cortex and subcortex. Furthermore, the correlation with functional connectivity was computed based on the components of tensor decomposition of communication score.
Benchmark and comparison with iOT-based simulation
Ideally, ligand receptor interactions should be validated by the spatial co-localization of the corresponding proteins, but this information is always lacking, especially at the whole brain scale. Therefore, we built a simulation method based on the iOT problem105,106,107. Actually, given the forward problem as a map \(\Phi :\left(M,L,R\right)\to \left(A,B,C\right)\), then the data synthesis is just the inverse map \({\Phi }^{-1}:\left(A,B,C\right)\to (M,L,R)\). Moreover, when \(\left(A,B,C\right)\) is observed without any noise, \(L,R\) are completely determined by \(\left(A,B,C\right)\). Thus, the inverse map reduces to \({\Phi }^{-1}:\left(A,B,C\right)\to M\). However, this map is not well-defined since transport cost \(M\) is not uniquely determined by \(A,B,C\) (e.g., any two transport costs differ by an additive constant that outputs the same optimal plan)107. Most existing papers focused on obtaining a unique approximation by placing constraints on the transport cost105,108,109. Following the same idea, we added two constraints on transport cost: (1) forcing the sum of all element to a given positive constant \(K\) (104 as default), and (2) adding an entropy term to the objective function. Therefore, this problem can be formulated as:
$$\begin{array}{c}\min {\left\langle M,A{{\mbox{diag}}}\left({C}^{{{\rm{T}}}}{{\mathbb{1}}}_{\left|I\right|}\right){B}^{{{\rm{T}}}}\right\rangle }_{F}-\varepsilon {\mbox {H}}(M)\\ {{\mbox{s.t.}}}{\sum }_{{{\rm{ij}}}}{M}_{{{\rm{ij}}}}=K\end{array}$$
Furthermore, this problem has a closed form solution. Let \(P=A{\mbox{diag}}\left({C}^{T}{{\mathbb{1}}}_{\left|I\right|}\right){B}^{T}\), we have
$$M=\frac{K\exp \left(-P/\varepsilon \right)}{{\sum }_{{ij}}{\left(\exp \left(-P/\varepsilon \right)\right)}_{{ij}}}$$
Synthetic data generation
There are two main steps for synthetic data: (1) generating a transport plan following the process that is commonly used in estimating a nonnegative tensor decomposition algorithm and (2) solving an iOT problem to get the transport cost. Specifically, the elements of \(A,B,C\) were generated randomly in interval \([0,\,1)\), and column normalization was performed on A and B. The ligand and receptor expression matrices are the sums along the first and second axes of the transport plan, which can be constructed by a factor matrix. Finally, the optimal value of the above iOT problem is the transport cost, and its low-rank representation can be solved using nonnegative matrix factorization. The CLRIA algorithm was compared with brain region N chosen from \(\left\{123,\,180\right\}\), number of LR-pairs chosen from \(\left\{300,\,500\right\}\), and column number of factor matrix r chosen from\(\left\{1,\,5,\,10,\,20,\,30\right\}\). For each case, ten independent simulations were carried out. The parameters of the entropy term and the marginal mass constraint were chosen from \(\{{0,\,10}^{-4},{10}^{-2}\}\) and \(\left\{1,\,2,\,5\right\}\). The low-rank constraint parameter was set the same as the r. The iteration stopped when both the relative error and the absolute error of the objective function was less than 10−5 with at least 300 iterations. For run-time comparisons, the data were simulated with brain regions in \(\left\{20,\,50,\,100,\,200,\,500,\,1000,\,2000\right\}\) with a fixed number of LR-pairs \(\left|I\right|=20\) and a column number \(r=5\) for efficiency.
Benchmark and comparison
Considering the permutation and scaling indeterminacy of the factor matrix110, we used CorrIndex111 (implemented in Tensorly112, version 0.8.1, to assess the consistency between CLRIA solutions and true values as well as to compare accuracy with other methods. The CorrIndex overcame these indeterminacies and showed its reliability and computational efficiency both theoretically and experimentally compared with other indices111. This index lies between 0 and 1, with a higher score indicating more dissimilarity. Because there is no method that can directly solve the factor matrix from the transport cost matrix (M) and the ligand and receptor expression matrices (L, R), we separated this process into two steps: (1) solving the transport plan and (2) decomposing the nonnegative tensor of the transport plan. The transport plan was solved using unbalanced OT (sinkhorn41 or MM algorithm113) and partial OT114 implemented in POT115 (version 0.9.1, as well as by the collective OT19. Then, the nonnegative CP decomposition was performed by the MU or HALS algorithm, which are classical NTD algorithms implemented in Tensorly112.
Brain network communication model
A communication model describes a strategy for guiding signal propagation along structural connectivity1,2. In this paper, we mainly focused on the transmission cost, which is defined as the mean first passage time (MFPT) on a biased random walk47. Specifically, a weighted connectome can be expressed as a matrix \(W\in {{\mathbb{R}}}^{N\times N}\), where \({W}_{{ij}}\) is the weighted streamline count. The weight-to-length transformation can be defined in various ways. Following a previous work28, we used \(D=-{\log }_{10}\left(W/\left(\max \left(W\right)+\min \left({W}_{ > 0}\right)\right)\right)\) to attenuate extreme weights70. The transmission cost was implemented based on the following dynamics of the system:
$${P}_{\lambda }\left(Y=j{{{\rm{| }}}}X=i,\,T=t\right)=\frac{1}{{Z}_{i}^{t}}\exp \left(-(\lambda \left({d}_{{ij}}+{g}_{{jt}}\right)+{d}_{{ij}})\right)$$
where X, Y and T are random variables indicating the current, next and target node of Markov chain, \({Z}_{i}^{t}={\sum }_{j}\exp (-(\lambda ({d}_{{ij}}+{g}_{{jt}})+{d}_{{ij}}))\) is a normalization factor, \({g}_{{jt}}\) is the shortest path length from j to t. Let transmission cost from i to t be \({C}_{\lambda }^{{trans}}(i,t)\), then we have
$${C}_{\lambda }^{{trans}}\left(i,t\right)={\sum }_{k}{n}_{\lambda }^{t}\left(i,k\right){c}_{\lambda }^{{trans}}\left(k,t\right)$$
where \({n}_{\lambda }^{t}\left(i,{k}\right)\) is the mean number of times that node k was visited by a walker starting at node i, and \({c}_{\lambda }^{{trans}}\left(k,t\right)={\sum }_{j}{P}_{\lambda }\left(Y=j| X=k,{T}=t\right){d}_{{kj}}\) is the expected distance that a walker at node i travels to its neighbor. Further computational details can be found in ref. 47. To construct a lambda spectrum of transmission costs representing changes from diffusion to routing strategy, thirty different \(\lambda\) values evenly spaced from −3 to 1.6 on a log scale were selected. To get rid of the magnitude of the transmission costs between different λ values, we rescaled the sum of all elements to 104.
Null model
For each λ value, we used a spatially constrained, label-permutation null model that randomly permutes region labels while preserving their spatial embedding and autocorrelation47. This method created a surface-based representation of cortical parcellations in fsaverage space. The fsaverage surface was then projected onto a sphere to define the spatial coordinates for each parcel. The vertices on the sphere were assigned on the basis of the shortest distance to the center of mass of each parcel. These parcel coordinates were then randomly rotated and the original parcels were reassigned the value of the closest rotated parcel. Of note, this procedure was performed at parcel resolution rather than vertex resolution to ensure that the parcel sizes were not changed during the rotation and reassignment procedure and was performed only for the left cortex. This method is available in the stats module of the netneurotools Python package ( For the subcortical region, random shuffles were performed following the ENIGMA toolbox116 (https://enigma-toolbox.readthedocs.io/en/latest/pages/08.spintest/index.html).
Asymmetric and trans-hierarchical signaling
The measures for the signaling asymmetry were inspired by Seguin et al.28. Specifically, given the transport plan \({{{\mathcal{P}}}}{{{\mathscr{\in }}}}{{\mathbb{R}}}^{N\times N\times \left|I\right|}\) reconstructed from tensor factor \(A,B,C\), where \({{{{\mathcal{P}}}}}_{{ijk}}\) denotes the communication strength between brain regions i and j mediated by the kth LR-pair, the difference in signaling flow between regions i and j can be defined as \({\Delta }_{{ijk}}={{{{\mathcal{P}}}}}_{{ijk}}-{{{{\mathcal{P}}}}}_{{jik}}\). We performed a one sample T-test to determine whether the mean of \({\Delta }_{{ijk}}\) across k was significantly different from 0, and the resulting T-statistics were used to quantify the extent of communication asymmetry. Repeating this test independently for all pairs of brain regions yielded a antisymmetric matrix \(T\in {{\mathbb{R}}}^{N\times N}\). A Bonferroni correction was performed to control for \(N\left(N-1\right)/2\) multiple comparisons and resulting P-values below 0.05 were considered as significantly asymmetric.
We used three definitions of cortical hierarchy to explore trans-hierarchy signaling: (1) anatomical hierarchy, which is defined by the T1w/T2w ratio to index the cortical microstructure and myelin content; (2) functional hierarchy, which is the gradient of functional connectivity52 involving the unimodal cortex at one end and the transmodal cortex at the other end; and (3) the sensorimotor-association axis (SA axis), which spans from the primary visual and somatomotor cortices to the transmodal association cortices and captures the spatial and temporal patterning of cortical maturation proceeds throughout childhood and adolescence54,55. All these macroscale maps were obtained from the neuromaps package117 in fsLR space and then aggregated into the Brainnetome parcellation. We compared the distribution of T-statistics between low-to-high and high-to-low asymmetric communication by the Kolmogorov-Smirnov test and spin test.
Energy spectral density of LR-pairs
To investigate the asymmetric signaling of each LR-pair, we used the difference between the outgoing and incoming degrees of the signaling network of the kth LR-pair as a measure. Specifically, let \({{{{\mathcal{P}}}}}_{k}\in {{\mathbb{R}}}^{N\times N}\) be the signaling network, then the difference \({d}_{k}={{{{\mathcal{P}}}}}_{k}{{\mathbb{1}}}_{N}-{{{{\mathcal{P}}}}}_{k}^{T}{{\mathbb{1}}}_{N}\in {{\mathbb{R}}}^{N}\), which can be viewed as a graph signal on the structure connectome. Following Preti et al.64, we defined the graph Fourier transformation (GFT) by the eigen decomposition of the graph Laplacian L, where L is the Laplacian transformation of the normalized structure connectivity. The eigenvalues \({\lambda }_{i}\) can be interpreted as frequencies, and the eigenmodes \({\xi }_{i}\) as structural connectome harmonics. Eigenmodes with a low eigenvalue encode smooth signals with respect to the structural network. Let \(U=\left[{\xi }_{1},\ldots,{\xi }_{N}\right]\in {{\mathbb{R}}}^{N\times N}\), the GFT converts a graph signal \({d}_{k}\) into its spectral representation \({\hat{d}}_{k}={U}^{T}{d}_{k}\), and its energy spectral density is defined as the square of \({\hat{d}}_{k}\).
Nonnegative tensor component analysis
We used the LR-pair loadings to identify the biological process by ClusterProfiler (version 4.7.1.001). Before performing the analysis, the LR-pair loadings were normalized across communication patterns to [0, 1] to remove the effect of LR-pairs expression. The genes involved with the top 20% of LR-pairs were defined as factor-related genes. The background genes were defined as all the ligand or receptor genes expressed in the brain. Terms with P-values below 0.05 after adjustment was considered to be significantly enriched functional terms.
We used the row-wise maximum of the factor matrix \(A,B\) to link a brain region to each factor. The ratio of these factor-specific regions in each intrinsic function network118 were denoted as \(p\). To further depict the specificity of a factor to the intrinsic function network, we defined a normalized entropy:
$$E\left(p\right)=-{\sum }_{i}{p}_{i}\log {p}_{i}/\log k$$
where k=8 is the number of intrinsic function network. \(\log k\) is a normalized factor to rescale the entropy to [0,1].
To further investigate the related cognitive function, a Neurosynth meta-analysis ( similar to the one implemented by Margulies et al.52 was conducted to assess topic terms associated with each factor. Values greater than 0.05 after row-sum normalization were defined as the ROI for each factor.
UOT regression
Given the ith columns of A and B, we generated a factor-specific communication pattern as their outer product and denoted it as \({X}_{i}\in {{\mathbb{R}}}^{N\times N}\). To investigate the brain state transition mediated by these communication patterns, we formulated it as an UOT regression problem. Specifically, if \(a,b\in {{\mathbb{R}}}^{N}\) are two brain states that represent stationary active patterns during the execution of cognitive processes, we aimed to minimize their transport distance:
$${\min }_{\beta,P}\;f \, \left(\beta,P\right)= {\left\langle M\left(\beta \right),P\right\rangle }_{F}-\varepsilon {\mbox {H}}\left(P\right)+{\lambda }_{1}{{\mbox{KL}}}\left(P{{\mathbb{1}}}_{N}\parallel a\right) \\ + {\lambda }_{2}{{\mbox{KL}}}\left({P}^{T}{{\mathbb{1}}}_{N}\parallel b\right)+\frac{\lambda }{2}\parallel \beta {\parallel }_{2}^{2}$$
where \(M\left(\beta \right)=\exp \left(-{\sum }_{i=1}^{r}{X}_{i}{\beta }_{i}\right),\beta \in {{\mathbb{R}}}^{r}\) can be viewed as the convex combination of the communication pattern with an affinity-to-distance transform. Following balanced optimal transport regression40, an alternative direction multiplier method was used to solve the problem, which mainly involved the following two steps:
$$\begin{array}{ccc}{P}^{(k+1)} &=& {{{{\rm{argmin}}}}}_{P}f\left({\beta }^{\left(k\right)},P\right)\\ {\beta }^{\left(k+1\right)} &=& {{{{\rm{argmin}}}}}_{\beta }f\left(\beta,{P}^{\left(k+1\right)}\right)\end{array}$$
In fact, we can obtain \({P}^{\left(k+1\right)}\) by the generalized Sinkhorn algorithm41 since it is the same as an entropic regularization optimal transport problem. By setting \(\frac{\partial f\left(\beta,P\right)}{\partial {\beta }_{i}}=0\), we have
$${\beta }_{i}=\frac{1}{\lambda }{{\mbox{tr}}}\left({{{\rm{X}}}}_{{{\rm{i}}}}\left(P\odot M\left(\beta \right)\right)\right),\,i=1,\ldots,r$$
Therefore, \(\beta\) can be solved by fixed point iteration and have nonnegativity. If \({T}_{i}\left(\beta \right)=\frac{1}{\lambda }{\mbox{tr}}\left({X}_{i}\left(P\odot M\left(\beta \right)\right)\right)\), it can be proved that \({T}_{i}\) is contractive mapping when \(\lambda \ge {\sum }_{{ij}}{P}_{{ij}}\) (not a tight bound), and thus we can conclude that \({T}_{i}\) is linearly convergent to the unique fixed point according to the Banach fixed-point theorem.
In practice, we set \(\varepsilon={\lambda }_{1}={\lambda }_{2}=1\), \(\lambda=2\) and compared the individual energy cost between 0 back to 2 back and 2 back to 0 back, which was measured as the UOT distance \({\left\langle M\left(\beta \right),P\right\rangle }_{F}\) and reflected the flexibility of the brain state. The significances were assessed by a repeat measures ANOVA with sex and age as covariates. To explore the communication pattern preference during these forward and reverse transition processes, the difference of \(\beta\) during the process can be used as a measure. To further investigate the underlying difference in the LR-pairs, we defined the \(C\beta\) as the importance of the LR-pairs during the transition for each individual. The significance was assessed by paired t test.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
link